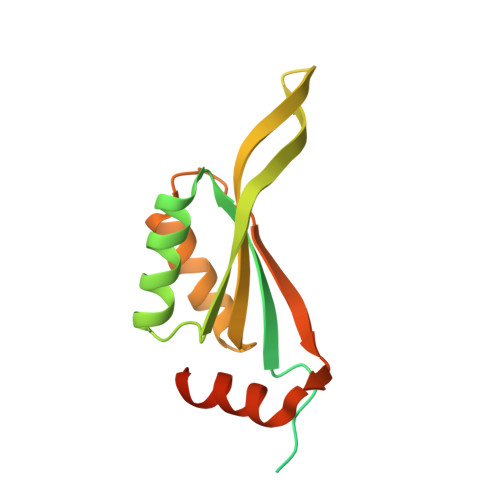
Structure of putative CutA1 from Homo sapiens determined at 2.05 A resolution.
Bagautdinov, B., Matsuura, Y., Bagautdinova, S., Kunishima, N., Yutani, K.(2008) Acta Crystallogr Sect F Struct Biol Cryst Commun 64: 351-357
- PubMed: 18453701 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309108009846
- Primary Citation Related Structures:
2ZFH - PubMed Abstract:
The structure of human brain CutA1 (HsCutA1) has been determined using diffraction data to 2.05 A resolution. HsCutA1 has been implicated in the anchoring of acetylcholinesterase in neuronal cell membranes, while its bacterial homologue Escherichia coli CutA1 is involved in copper tolerance. Additionally, the structure of HsCutA1 bears similarity to that of the signal transduction protein PII, which is involved in regulation of nitrogen metabolism. Although several crystal structures of CutA1 from various sources with different rotation angles and degrees of interaction between trimer interfaces have been reported, the specific functional role of CutA1 is still unclear. In this study, the X-ray structure of HsCutA1 was determined in space group P2(1)2(1)2(1), with unit-cell parameters a = 68.69, b = 88.84, c = 125.33 A and six molecules per asymmetric unit. HsCutA1 is a trimeric molecule with intertwined antiparallel beta-strands; each subunit has a molecular weight of 14.6 kDa and contains 135 amino-acid residues. In order to obtain clues to the possible function of HsCutA1, its crystal structure was compared with those of other CutA1 and PII proteins.
- Protein Structure Analysis Team, RIKEN SPring-8 Center, Harima Institute, 1-1-1 Kouto, Sayo-cho, Sayo-gun, Hyogo 679-5148, Japan. bagautdi@spring8.or.jp
Organizational Affiliation: