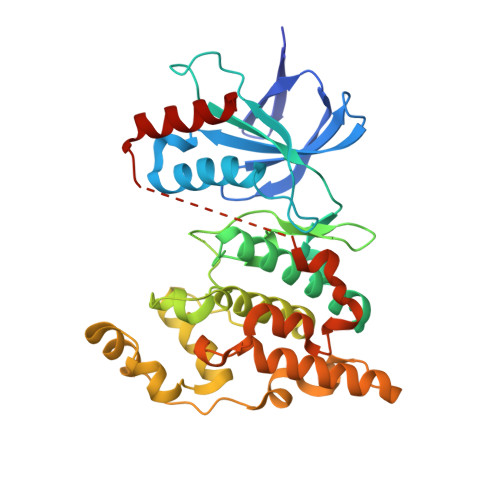
Discovery, synthesis and biological evaluation of isoquinolones as novel and highly selective JNK inhibitors (1)
Asano, Y., Kitamura, S., Ohra, T., Aso, K., Igata, H., Tamura, T., Kawamoto, T., Tanaka, T., Sogabe, S., Matsumoto, S., Yamaguchi, M., Kimura, H., Itoh, F.(2008) Bioorg Med Chem 16: 4715-4732
- PubMed: 18313304 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmc.2008.02.027
- Primary Citation Related Structures:
2ZDU - PubMed Abstract:
A novel series of 4-phenylisoquinolones were synthesized and evaluated as c-Jun N-terminal kinase (JNK) inhibitors. Initial modification at the 2- and 3-positions of the isoquinolone ring of hit compound 4, identified from high-throughput screening, led to the lead compound 6b. The optimization was carried out using a JNK1-binding model of 6b and several compounds exhibited potent JNK inhibition. Among them, 11g significantly inhibited cardiac hypertrophy in rat pressure-overload models without affecting blood pressure and the concept of JNK inhibitors as novel therapeutic agents for heart failure was confirmed.
- Medicinal Chemistry Research Laboratories, Pharmaceutical Research Division, Takeda Pharmaceutical Company, Ltd, 17-85, Jusohonmachi 2-chome, Osaka 532-8686, Japan. Asano_Yasutomi@takeda.co.jp
Organizational Affiliation: