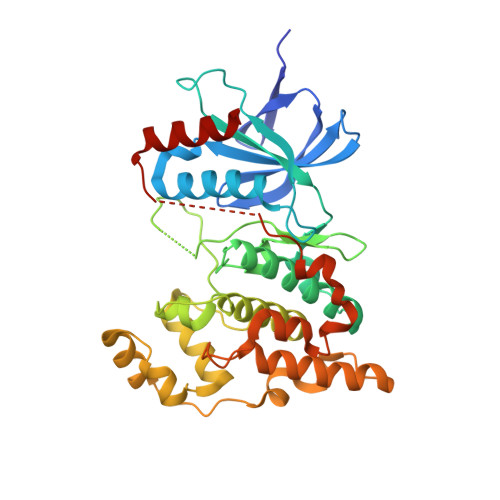
Discovery, synthesis and biological evaluation of isoquinolones as novel and highly selective JNK inhibitors (2)
Asano, Y., Kitamura, S., Ohra, T., Itoh, F., Kajino, M., Tamura, T., Kaneko, M., Ikeda, S., Igata, H., Kawamoto, T., Sogabe, S., Matsumoto, S., Tanaka, T., Yamaguchi, M., Kimura, H., Fukumoto, S.(2008) Bioorg Med Chem 16: 4699-4714
- PubMed: 18313930 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmc.2008.02.028
- Primary Citation Related Structures:
2ZDT - PubMed Abstract:
3-Metoxycarbonyl isoquinolone derivative 1 has been identified as a potent JNK inhibitor and significantly inhibited cardiac hypertrophy in a rat pressure-overload model. Herein, a series of isoquinolones with an imidazolylmethyl or a pyrazolylmethyl group at the 2-position were designed based on X-ray crystallographic analysis of the complex between the isoquinolone compound and JNK3, as wells as the relationship between compound lipophilicity (logD) and activity in a cell-based assay. The compounds prepared showed potent JNK1 inhibitory activities in a cell-based assay. Among them the isoquinolone derivative possessing 5-[(cyclopropylamino)carbonyl]-1-methyl-1H-pyrazole (16e) exhibited significant anti-hypertrophic activity at doses of more than 1mg/kg (po) in a pressure-overload model.
- Medicinal Chemistry Research Laboratories, Pharmaceutical Research Division, Takeda Pharmaceutical Company, Ltd, 17-85, Jusohonmachi 2-chome, Osaka 532-8686, Japan. Asano_Yasutimi@takeda.co.jp
Organizational Affiliation: