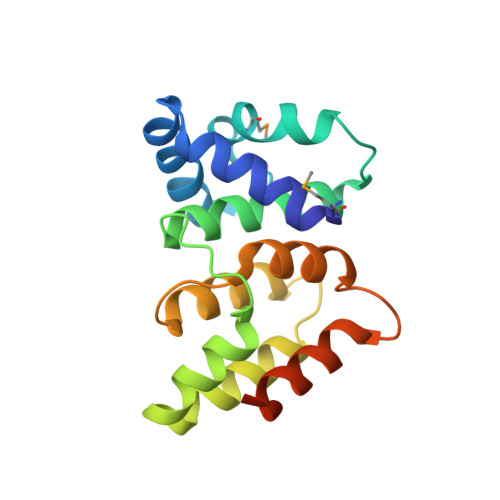
X-ray crystal structure of a CRISPR-associated protein, Cse2, from Thermus thermophilus HB8
Agari, Y., Yokoyama, S., Kuramitsu, S., Shinkai, A.(2008) Proteins 73: 1063-1067
- PubMed: 18798563
- DOI: https://doi.org/10.1002/prot.22224
- Primary Citation of Related Structures:
2ZCA - Photon Science Research Division, RIKEN SPring-8 Center, Harima Institute, 1-1-1 Kouto, Sayo, Hyogo 679-5148, Japan.
Organizational Affiliation: