The crystal structure of the all cysteinyl coordinated D14C variant of [4Fe-4S] Pyrococcus furiosus ferredoxin
Johannessen, M.N., Nielsen, M.S., Ooi, B.L., Christensen, H.E.M., Harris, P.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Ferredoxin | 66 | Pyrococcus furiosus DSM 3638 | Mutation(s): 1 Gene Names: fdxA | 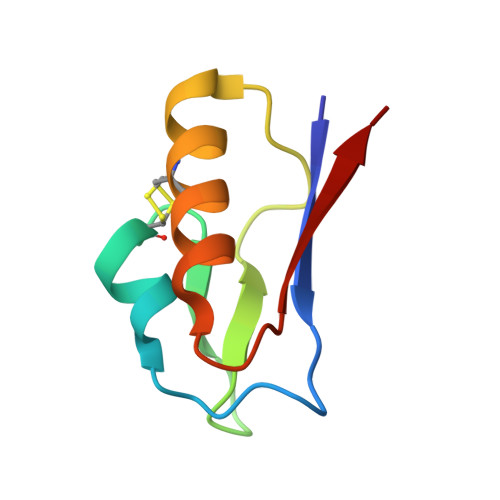 | |
UniProt | |||||
Find proteins for P29603 (Pyrococcus furiosus (strain ATCC 43587 / DSM 3638 / JCM 8422 / Vc1)) Explore P29603 Go to UniProtKB: P29603 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P29603 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| SF4 Query on SF4 | C [auth A], F [auth B] | IRON/SULFUR CLUSTER Fe4 S4 LJBDFODJNLIPKO-UHFFFAOYSA-N |  | ||
| NCO Query on NCO | D [auth A], E [auth A], G [auth B], H [auth B] | COBALT HEXAMMINE(III) Co H18 N6 DYLMFCCYOUSRTK-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 51.4 | α = 90 |
| b = 116.8 | β = 90 |
| c = 47.7 | γ = 90 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| ProDC | data collection |
| XDS | data reduction |
| XSCALE | data scaling |
| MOLREP | phasing |