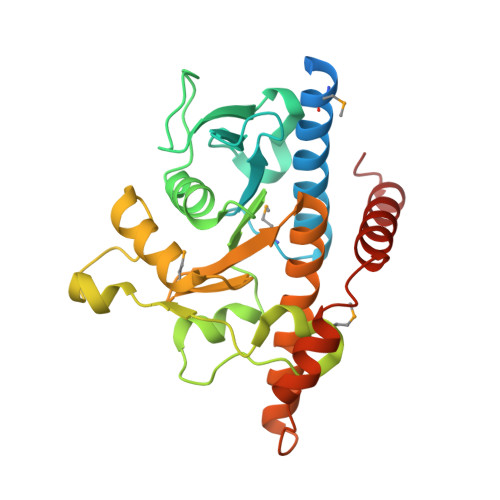
Gene Identification and Structural Characterization of the Pyridoxal 5'-Phosphate Degradative Protein 3-Hydroxy-2-methylpyridine-4,5-dicarboxylate Decarboxylase from Mesorhizobium loti MAFF303099
Mukherjee, T., McCulloch, K.M., Ealick, S.E., Begley, T.P.(2007) Biochemistry 46: 13606-13615
- PubMed: 17973403
- DOI: https://doi.org/10.1021/bi701439j
- Primary Citation Related Structures:
2Z7B - PubMed Abstract:
The function of the mlr6791 gene from Mesorhizobium loti MAFF303099 has been identified. This gene encodes 3-hydroxy-2-methylpyridine-4,5-dicarboxylate decarboxylase (HMPDdc), an enzyme involved in the catabolism of pyridoxal 5'-phosphate (Vitamin B6). This enzyme was overexpressed in Escherichia coli and characterized. HMPDdc is a 26 kDa protein that catalyzes the decarboxylation of 3-hydroxy-2-methylpyridine-4,5-dicarboxylate to 3-hydroxy-2-methylpyridine-5-carboxylate. The KM and kcat were found to be 366 microM and 0.6 s-1, respectively. The structure of this enzyme was determined at 1.9 A resolution using SAD phasing and belongs to the class II aldolase/adducin superfamily. While the decarboxylation of hydroxy-substituted benzene rings is a common motif in biosynthesis, the mechanism of this reaction is still poorly characterized. The structural studies described here suggest that catalysis of such decarboxylations proceeds by an aldolase-like mechanism.
- Department of Chemistry and Chemical Biology, Cornell University, Ithaca, New York 14853, USA.
Organizational Affiliation: