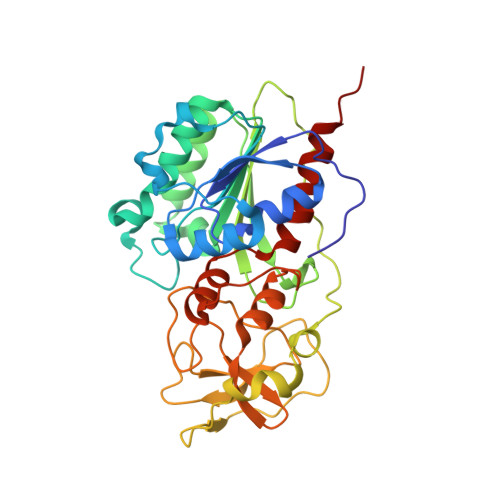
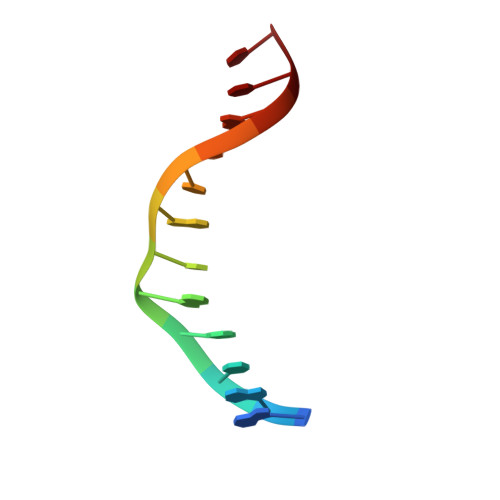
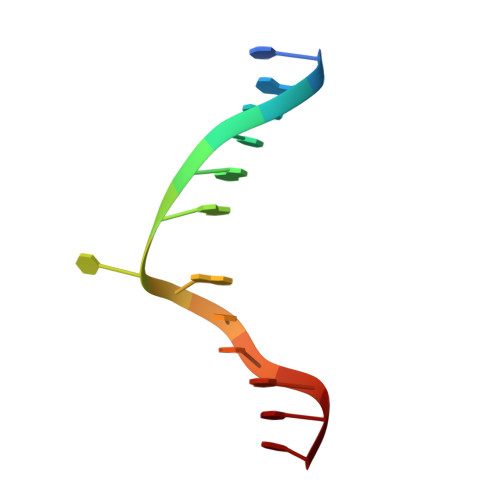
S-Adenosyl-l-methionine-Dependent Methyl Transfer: Observable Precatalytic Intermediates during DNA Cytosine Methylation
Youngblood, B., Shieh, F.K., Buller, F., Bullock, T., Reich, N.O.(2007) Biochemistry 46: 8766-8775
- PubMed: 17616174
- DOI: https://doi.org/10.1021/bi7005948
- Primary Citation of Related Structures:
2Z6A - PubMed Abstract:
S-adenosyl-L-methionine- (AdoMet-) dependent methyltransferases are widespread, play critical roles in diverse biological pathways, and are antibiotic and cancer drug targets. Presently missing from our understanding of any AdoMet-dependent methyl-transfer reaction is a high-resolution structure of a precatalytic enzyme/AdoMet/DNA complex. The catalytic mechanism of DNA cytosine methylation was studied by structurally and functionally characterizing several active site mutants of the bacterial enzyme M.HhaI. The 2.64 A resolution protein/DNA/AdoMet structure of the inactive C81A M.HhaI mutant suggests that active site water, an approximately 13 degree tilt of the target base toward the active site nucleophile, and the presence or absence of the cofactor methylsulfonium are coupled via a hydrogen-bonding network involving Tyr167. The active site in the mutant complex is assembled to optimally align the pyrimidine for nucleophilic attack and subsequent methyl transfer, consistent with previous molecular dynamics ab initio and quantum mechanics/molecular mechanics calculations. The mutant/DNA/AdoHcy structure (2.88 A resolution) provides a direct comparison to the postcatalytic complex. A third C81A ternary structure (2.22 A resolution) reveals hydrolysis of AdoMet to adenosine in the active site, further validating the coupling between the methionine portion of AdoMet and ultimately validating the structural observation of a prechemistry/postchemistry water network. Disruption of this hydrogen-bonding network by a Tyr167 to Phe167 mutation does not alter the kinetics of nucleophilic attack or methyl transfer. However, the Y167F mutant shows detectable changes in kcat, caused by the perturbed kinetics of AdoHcy release. These results provide a basis for including an extensive hydrogen-bonding network in controlling the rate-limiting product release steps during cytosine methylation.
- Program in Biomolecular Science and Engineering, University of California, Santa Barbara, California 93106-9510, USA.
Organizational Affiliation: