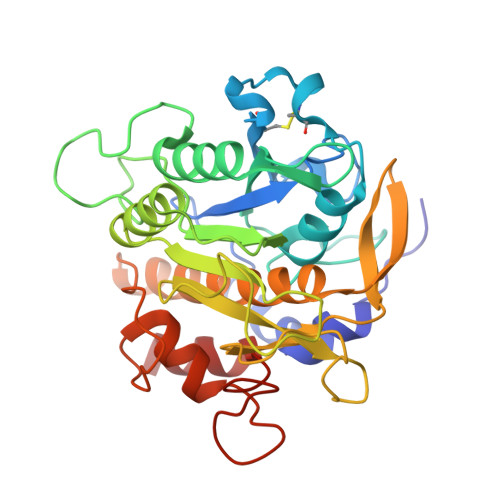
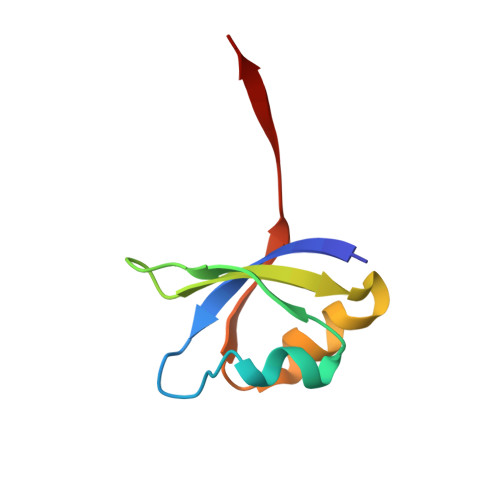
Requirement of left-handed glycine residue for high stability of the Tk-subtilisin propeptide as revealed by mutational and crystallographic analyses
Pulido, M.A., Tanaka, S., Sringiew, C., You, D.J., Matsumura, H., Koga, Y., Takano, K., Kanaya, S.(2007) J Mol Biology 374: 1359-1373
- PubMed: 17988685
- DOI: https://doi.org/10.1016/j.jmb.2007.10.030
- Primary Citation of Related Structures:
2Z56, 2Z57, 2Z58 - PubMed Abstract:
Tk-subtilisin [the mature domain of Pro-Tk-subtilisin in active form (Gly70-Gly398)] from the hyperthermophilic archaeon Thermococcus kodakaraensis is matured from Pro-Tk-subtilisin [a subtilisin homologue from T. kodakaraensis in pro form (Gly1-Gly398)] upon autoprocessing and degradation of propeptide. Pro-Tk-subtilisin is characterized by extremely slow maturation at mild temperatures, but this maturation rate is greatly increased by a single Gly56-->Ser mutation in the propeptide region. To analyze the role of Gly56, which assumes a left-handed conformation, Pro-Tk-subtilisin variants with complete amino acid substitutions at Gly56 were constructed. A comparison of their halo-forming activities suggests that all variants, except for Pro-G56W [Pro-G56X, Pro-Tk-subtilisin with Gly56-->X mutation (X = any amino acid)], mature faster than WT. Pro-G56W and Pro-G56E with the lowest and highest maturation rates, respectively, among 19 variants, as well as WT and Pro-G56S, were overproduced, purified, and characterized. SDS-PAGE analyses and Tk-subtilisin activity assay indicated that their maturation rates increased in the order WT < or = Pro-G56W < Pro-G56S < Pro-G56E. The propeptides of these variants were also overproduced, purified, and characterized. The stability and inhibitory potency of these propeptides decreased in the order Tk-propeptide [propeptide of Tk-subtilisin (Gly1-Leu69)] > or = G56W-propeptide > G56S-propeptide > G56E-propeptide, indicating that they are inversely correlated with the maturation rates of Pro7-Tk-subtilisin and its derivatives. The crystal structures of these propeptides determined in complex with S324A-subtilisin indicate that the conformation of the propeptide is altered by the mutation, such that nonglycine residues at position 56 assume a right-handed conformation and hydrophobic interactions at the core region decrease. These results indicate that Gly56 is required in stabilizing the propeptide fold. Stabilization of this fold leads to strong binding of Tk-propeptide to Tk-subtilisin, high resistance of Tk-propeptide to proteolytic degradation, and slow maturation of Pro-Tk-subtilisin.
- Department of Material and Life Science, Graduate School of Engineering, Osaka University, 2-1 Yamadaoka, Suita, Osaka 565-0871, Japan.
Organizational Affiliation: