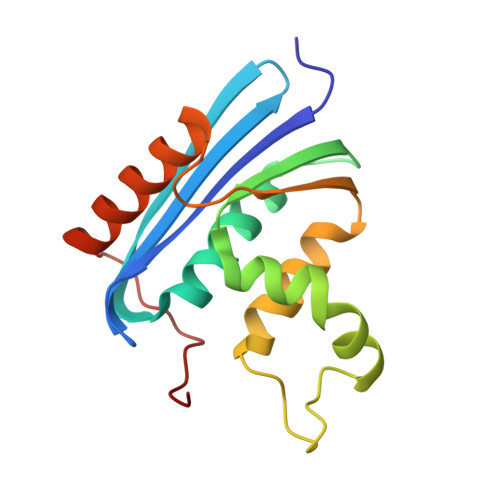
Structural and thermodynamic analyses of Escherichia coli RNase HI variant with quintuple thermostabilizing mutations.
Haruki, M., Tanaka, M., Motegi, T., Tadokoro, T., Koga, Y., Takano, K., Kanaya, S.(2007) FEBS J 274: 5815-5825
- PubMed: 17944939 Search on PubMed
- DOI: https://doi.org/10.1111/j.1742-4658.2007.06104.x
- Primary Citation Related Structures:
2YV0 - PubMed Abstract:
A combination of five thermostabilizing mutations, Gly23-->Ala, His62-->Pro, Val74-->Leu, Lys95-->Gly, and Asp134-->His, has been shown to additively enhance the thermostability of Escherichia coli RNase HI [Akasako A, Haruki M, Oobatake M & Kanaya S (1995) Biochemistry34, 8115-8122]. In this study, we determined the crystal structure of the protein with these mutations (5H-RNase HI) to analyze the effects of the mutations on the structure in detail. The structures of the mutation sites were almost identical to those of the mutant proteins to which the mutations were individually introduced, except for G23A, for which the structure of the single mutant protein is not available. Moreover, only slight changes in the backbone conformation of the protein were observed, and the interactions of the side chains were almost conserved. These results indicate that these mutations almost independently affect the protein structure, and are consistent with the fact that the thermostabiling effects of the mutations are cumulative. We also determined the protein stability curve describing the temperature dependence of the free energy of unfolding of 5H-RNase HI to elucidate the thermostabilization mechanism. The maximal stability for 5H-RNase HI was as high as that for the cysteine-free variant of Thermus thermophilus RNase HI. In contrast, the heat capacity of unfolding for 5H-RNase H was similar to that for E. coli RNase HI, which is considerably higher than that for T. thermophilus RNase HI. These results suggest that 5H-RNase HI is stabilized, in part, by the thermostabilization mechanism adopted by T. thermophilus RNase HI.
- Department of Materials Chemistry and Engineering, Nihon University, Koriyama, Japan. haruki@chem.ce.nihon-u.ac.jp
Organizational Affiliation: