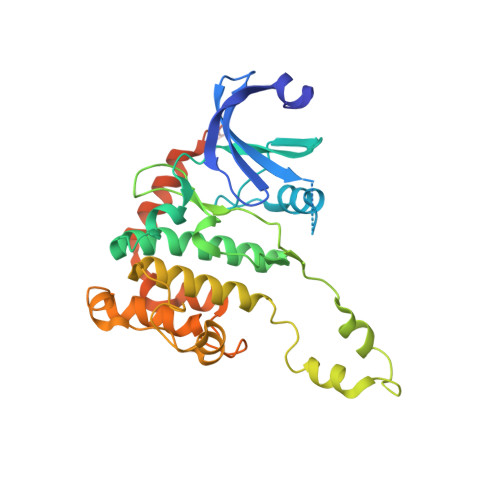
X-Ray Structures of Checkpoint Kinase 2 in Complex with Inhibitors that Target its Gatekeeper-Dependent Hydrophobic Pocket.
Lountos, G.T., Jobson, A.G., Tropea, J.E., Self, C.R., Zhang, G., Pommier, Y., Shoemaker, R.H., Waugh, D.S.(2011) FEBS Lett 585: 3245
- PubMed: 21907711 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.febslet.2011.08.050
- Primary Citation Related Structures:
2YIQ, 2YIR, 2YIT - PubMed Abstract:
The serine/threonine checkpoint kinase 2 (Chk2) is an attractive molecular target for the development of small molecule inhibitors to treat cancer. Here, we report the rational design of Chk2 inhibitors that target the gatekeeper-dependent hydrophobic pocket located behind the adenine-binding region of the ATP-binding site. These compounds exhibit IC(50) values in the low nanomolar range and are highly selective for Chk2 over Chk1. X-ray crystallography was used to determine the structures of the inhibitors in complex with the catalytic kinase domain of Chk2 to verify their modes of binding.
- Basic Science Program, SAIC-Frederick, Frederick, MD 21702-1201, USA.
Organizational Affiliation: