Surprising Complexity of the Asf1 Histone Chaperone-Rad53 Kinase Interaction
Jiao, Y., Seeger, K., Lautrette, A., Gaubert, A., Mousson, F., Guerois, R., Mann, C., Ochsenbein, F.(2012) Proc Natl Acad Sci U S A 109: 2866
- PubMed: 22323608 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1106023109
- Primary Citation Related Structures:
2YGV - PubMed Abstract:
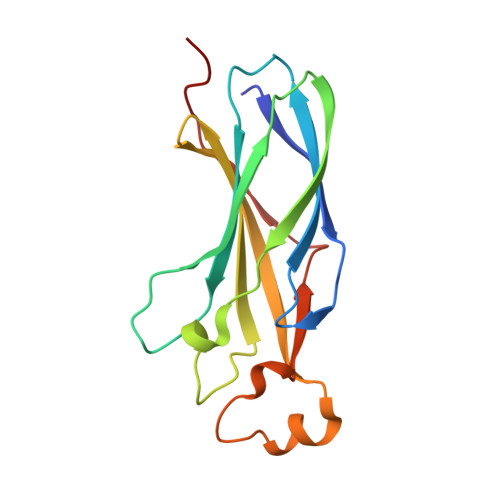
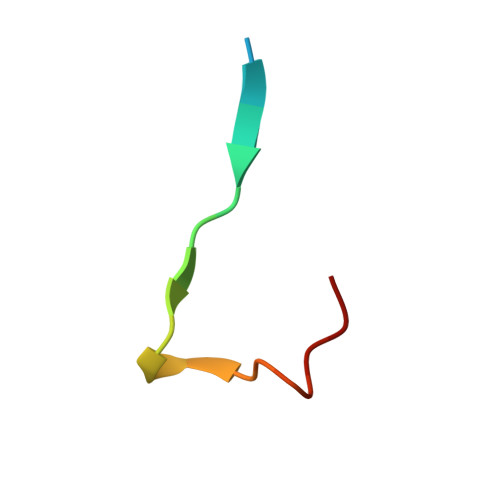
The histone chaperone Asf1 and the checkpoint kinase Rad53 are found in a complex in budding yeast cells in the absence of genotoxic stress. Our data suggest that this complex involves at least three interaction sites. One site involves the H3-binding surface of Asf11 with an as yet undefined surface of Rad53. A second site is formed by the Rad53-FHA1 domain binding to Asf1-T(270) phosphorylated by casein kinase II. The third site involves the C-terminal 21 amino acids of Rad53 bound to the conserved Asf1 N-terminal domain. The structure of this site showed that the Rad53 C-terminus binds Asf1 in a remarkably similar manner to peptides derived from the histone cochaperones HirA and CAF-I. We call this binding motif, (R/K)R(I/A/V) (L/P), the AIP box for Asf1-Interacting Protein box. Furthermore, C-terminal Rad53-F(820) binds the same pocket of Asf1 as does histone H4-F(100). Thus Rad53 competes with histones H3-H4 and cochaperones HirA/CAF-I for binding to Asf1. Rad53 is phosphorylated and activated upon genotoxic stress. The Asf1-Rad53 complex dissociated when cells were treated with hydroxyurea but not methyl-methane-sulfonate, suggesting a regulation of the complex as a function of the stress. We identified a rad53 mutation that destabilized the Asf1-Rad53 complex and increased the viability of rad9 and rad24 mutants in conditions of genotoxic stress, suggesting that complex stability impacts the DNA damage response.
- Commissariat à l'Energie Atomique, iBiTec-S, Service de Biologie Intégrative et de Génétique Moléculaire, Gif-sur-Yvette F-91191, France.
Organizational Affiliation: