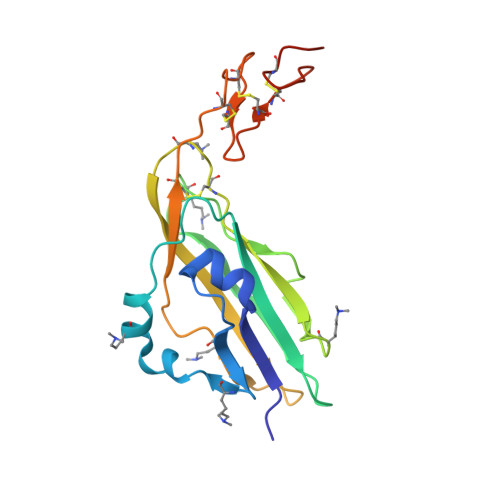
Modular Mechanism of Wnt Signaling Inhibition by Wnt Inhibitory Factor 1
Malinauskas, T., Aricescu, A.R., Lu, W., Siebold, C., Jones, E.Y.(2011) Nat Struct Mol Biol 18: 886
- PubMed: 21743455 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.2081
- Primary Citation Related Structures:
2YGN, 2YGO, 2YGP, 2YGQ - PubMed Abstract:
Wnt morphogens control embryonic development and homeostasis in adult tissues. In vertebrates the N-terminal WIF domain (WIF-1(WD)) of Wnt inhibitory factor 1 (WIF-1) binds Wnt ligands. Our crystal structure of WIF-1(WD) reveals a previously unidentified binding site for phospholipid; two acyl chains extend deep into the domain, and the head group is exposed to the surface. Biophysical and cellular assays indicate that there is a WIF-1(WD) Wnt-binding surface proximal to the lipid head group but also implicate the five epidermal growth factor (EGF)-like domains (EGFs I-V) in Wnt binding. The six-domain WIF-1 crystal structure shows that EGFs I-V are wrapped back, interfacing with WIF-1(WD) at EGF III. EGFs II-V contain a heparan sulfate proteoglycan (HSPG)-binding site, consistent with conserved positively charged residues on EGF IV. This combination of HSPG- and Wnt-binding properties suggests a modular model for the localization of WIF-1 and for signal inhibition within morphogen gradients.
- Division of Structural Biology, Wellcome Trust Centre for Human Genetics, University of Oxford, Oxford, UK.
Organizational Affiliation: