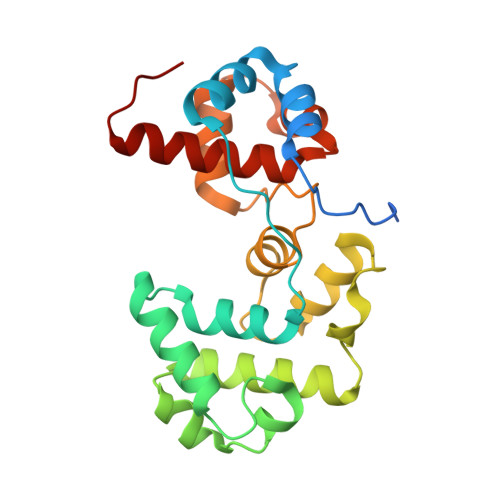
Structure-function studies of an unusual 3-methyladenine DNA glycosylase II (AlkA) from Deinococcus radiodurans.
Moe, E., Hall, D.R., Leiros, I., Monsen, V.T., Timmins, J., McSweeney, S.(2012) Acta Crystallogr D Biol Crystallogr 68: 703-712
- PubMed: 22683793 Search on PubMed
- DOI: https://doi.org/10.1107/S090744491200947X
- Primary Citation Related Structures:
2YG8, 2YG9 - PubMed Abstract:
3-Methyladenine DNA glycosylase II (AlkA) is a DNA-repair enzyme that removes alkylated bases in DNA via the base-excision repair (BER) pathway. The enzyme belongs to the helix-hairpin-helix (HhH) superfamily of DNA glycosylases and possesses broad substrate specificity. In the genome of Deinococcus radiodurans, two genes encoding putative AlkA have been identified (Dr_2074 and Dr_2584). Dr_2074 is a homologue of human AlkA (MPG or AAG) and Dr_2584 is a homologue of bacterial AlkAs. Here, the three-dimensional structure of Dr_2584 (DrAlkA2) is presented and compared with the previously determined structure of Escherichia coli AlkA (EcAlkA). The results show that the enzyme consists of two helical-bundle domains separated by a wide DNA-binding cleft and contains an HhH motif. Overall, the protein fold is similar to the two helical-bundle domains of EcAlkA, while the third N-terminal mixed α/β domain observed in EcAlkA is absent. Substrate-specificity analyses show that DrAlkA2, like EcAlkA, is able to remove both 3-methyladenine (3meA) and 7-methylguanine (7meG) from DNA; however, the enzyme possesses no activity towards 1,N(6)-ethenoadenine (ℇA) and hypoxanthine (Hx). In addition, it shows activity towards the AlkB dioxygenase substrates 3-methylcytosine (3meC) and 1-methyladenine (1meA). Thus, the enzyme seems to preferentially repair methylated bases with weakened N-glycosidic bonds; this is an unusual specificity for a bacterial AlkA protein and is probably dictated by a combination of the wide DNA-binding cleft and a highly accessible specificity pocket.
- The Norwegian Structural Biology Centre (NorStruct), Department of Chemistry, University of Tromsø, N-9037 Tromsø, Norway.
Organizational Affiliation: