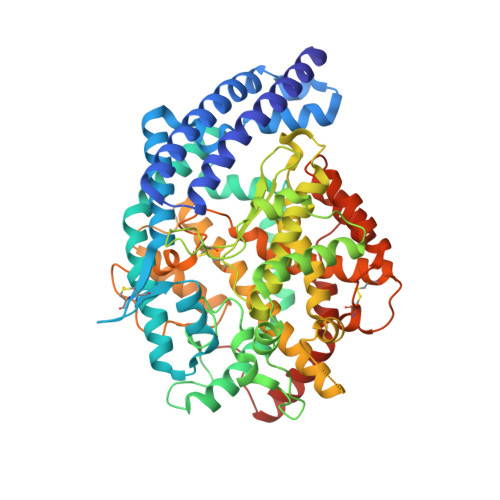
Structural Characterization of Angiotensin-I Converting Enzyme in Complex with a Selenium Analogue of Captopril
Akif, M., Masuyer, G., Schwager, S.L.U., Bhuyan, B.J., Mugesh, G., Isaac, R.E., Sturrock, E.D., Acharya, K.R.(2011) FEBS J 278: 3644
- PubMed: 21810173 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1111/j.1742-4658.2011.08276.x
- Primary Citation Related Structures:
2YDM, 3ZQZ - PubMed Abstract:
Human somatic angiotensin I-converting enzyme (ACE), a zinc-dependent dipeptidyl carboxypeptidase, is central to the regulation of the renin-angiotensin aldosterone system. It is a well-known target for combating hypertension and related cardiovascular diseases. In a recent study by Bhuyan and Mugesh [Org. Biomol. Chem. (2011) 9, 1356-1365], it was shown that the selenium analogues of captopril (a well-known clinical inhibitor of ACE) not only inhibit ACE, but also protect against peroxynitrite-mediated nitration of peptides and proteins. Here, we report the crystal structures of human testis ACE (tACE) and a homologue of ACE, known as AnCE, from Drosophila melanogaster in complex with the most promising selenium analogue of captopril (SeCap) determined at 2.4 and 2.35 Å resolution, respectively. The inhibitor binds at the active site of tACE and AnCE in an analogous fashion to that observed for captopril and provide the first examples of a protein-selenolate interaction. These new structures of tACE-SeCap and AnCE-SeCap inhibitor complexes presented here provide important information for further exploration of zinc coordinating selenium-based ACE inhibitor pharmacophores with significant antioxidant activity.
- Department of Biology and Biochemistry, University of Bath, Bath, UK.
Organizational Affiliation: