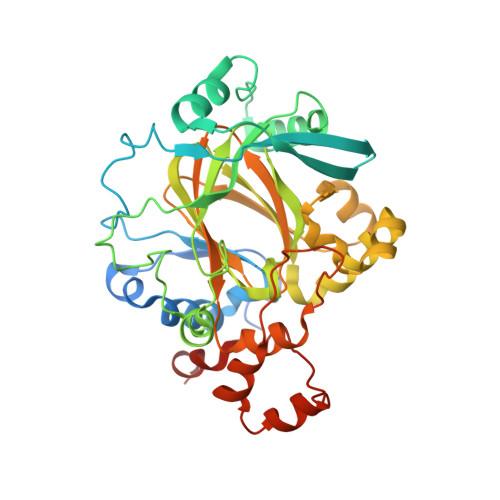
The oncometabolite 2-hydroxyglutarate inhibits histone lysine demethylases.
Chowdhury, R., Yeoh, K.K., Tian, Y.M., Hillringhaus, L., Bagg, E.A., Rose, N.R., Leung, I.K., Li, X.S., Woon, E.C., Yang, M., McDonough, M.A., King, O.N., Clifton, I.J., Klose, R.J., Claridge, T.D., Ratcliffe, P.J., Schofield, C.J., Kawamura, A.(2011) EMBO Rep 12: 463-469
- PubMed: 21460794 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/embor.2011.43
- Primary Citation Related Structures:
2YBK, 2YBP, 2YBS, 2YC0, 2YDE - PubMed Abstract:
Mutations in isocitrate dehydrogenases (IDHs) have a gain-of-function effect leading to R(-)-2-hydroxyglutarate (R-2HG) accumulation. By using biochemical, structural and cellular assays, we show that either or both R- and S-2HG inhibit 2-oxoglutarate (2OG)-dependent oxygenases with varying potencies. Half-maximal inhibitory concentration (IC(50)) values for the R-form of 2HG varied from approximately 25 μM for the histone N(ɛ)-lysine demethylase JMJD2A to more than 5 mM for the hypoxia-inducible factor (HIF) prolyl hydroxylase. The results indicate that candidate oncogenic pathways in IDH-associated malignancy should include those that are regulated by other 2OG oxygenases than HIF hydroxylases, in particular those involving the regulation of histone methylation.
- Chemistry Research Laboratory, Department of Chemistry, University of Oxford, Mansfield Road, Oxford OX1 3TA, UK.
Organizational Affiliation: