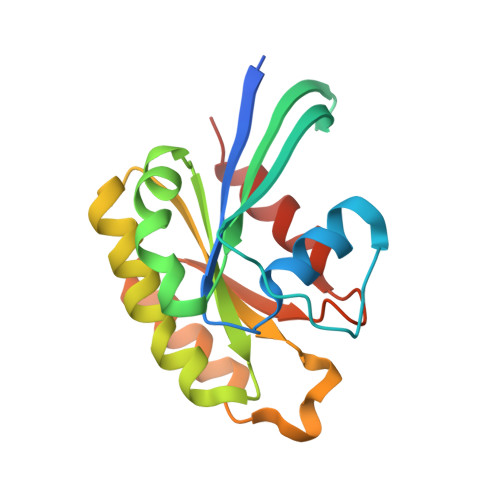
Structure of the Drosophila Melanogaster Rab6 Gtpase at 1.4 A Resolution
Walden, M., Jenkins, H.T., Edwards, T.A.(2011) Acta Crystallogr Sect F Struct Biol Cryst Commun 67: 744
- PubMed: 21795785 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309111017453
- Primary Citation Related Structures:
2Y8E - PubMed Abstract:
Rab6 is a small GTPase that belongs to the p21 Ras superfamily. It is involved in vesicle trafficking between the Golgi apparatus and endosomes/ER in eukaryotes. The GDP-bound inactive protein undergoes conformational changes when the nucleotide is exchanged to GTP, allowing Rab6 to interact with a variety of different effector proteins. To further understand how these changes affect downstream protein binding, the crystal structure of Rab6 from Drosophila melanogaster has been solved to 1.4 Å resolution, the highest resolution for a Rab6 structure to date. The crystals belonged to space group C2, with unit-cell parameters a=116.5, b=42.71, c=86.86 Å, α=90, β=133.12, γ=90°. The model was refined to an R factor of 14.5% and an Rfree of 17.3%.
- Astbury Centre for Structural Molecular Biology, University of Leeds, Leeds LS2 9JT, England.
Organizational Affiliation: