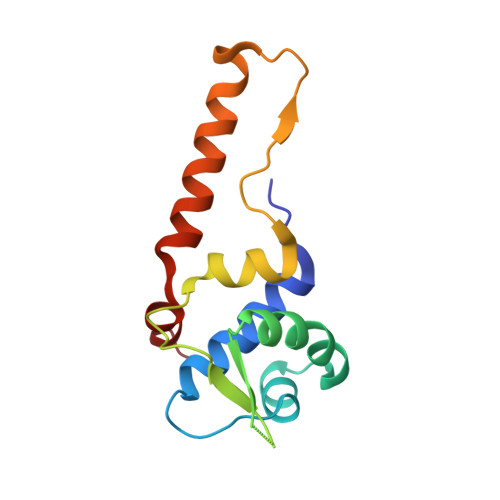
Insights Into the Rrf2 Repressor Family - the Structure of Cymr, the Global Cysteine Regulator of Bacillus Subtilis.
Shepard, W., Soutourina, O., Courtois, E., England, P., Haouz, A., Martin-Verstraete, I.(2011) FEBS J 278: 2689
- PubMed: 21624051
- DOI: https://doi.org/10.1111/j.1742-4658.2011.08195.x
- Primary Citation Related Structures:
2Y75 - PubMed Abstract:
The global regulator CymR represses the transcription of a large set of genes involved in cystine uptake and cysteine biosynthesis in Bacillus subtilis and Staphylococcus aureus. This repressor belongs to the widespread and poorly characterized Rrf2 family of regulators. The crystal structure of CymR from B. subtilis reveals a biologically active dimer, where each monomer folds into two tightly packed domains: a DNA-binding domain, which houses a winged helix-turn-helix (wHTH) motif; and a long dimerization domain, which places the wHTH motifs at the extremes. This architecture explains how these small regulators can span 23-27-bp DNA targets. The wHTH motif of CymR resembles those of the GntR superfamily of regulators, such as FadR and HutC. Superimposing the FadR wHTH motifs bound to their DNA fragments onto the wHTH motifs of the CymR dimer structure suggests that the DNA target and/or the protein must undergo some conformational changes upon binding. The CymR structure also hints at a possible location of the Fe-S centre associated with several Rrf2-type regulators.
- Synchrotron SOLEIL, L'Orme des Merisiers, Saint Aubin, BP48, Gif-sur-Yvette, France.
Organizational Affiliation: