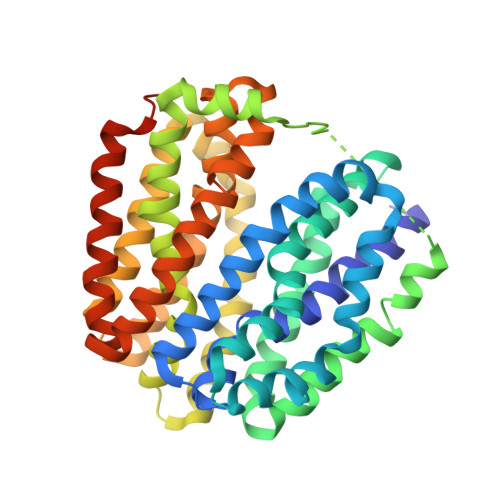
Crystal Structure of Lactose Permease in Complex with an Affinity Inactivator Yields Unique Insight Into Sugar Recognition.
Chaptal, V., Kwon, S., Sawaya, M.R., Guan, L., Kaback, H.R., Abramson, J.(2011) Proc Natl Acad Sci U S A 108: 9361
- PubMed: 21593407 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1105687108
- Primary Citation Related Structures:
2Y5Y - PubMed Abstract:
Lactose permease of Escherichia coli (LacY) with a single-Cys residue in place of A122 (helix IV) transports galactopyranosides and is specifically inactivated by methanethiosulfonyl-galactopyranosides (MTS-gal), which behave as unique suicide substrates. In order to study the mechanism of inactivation more precisely, we solved the structure of single-Cys122 LacY in complex with covalently bound MTS-gal. This structure exhibits an inward-facing conformation similar to that observed previously with a slight narrowing of the cytoplasmic cavity. MTS-gal is bound covalently, forming a disulfide bond with C122 and positioned between R144 and W151. E269, a residue essential for binding, coordinates the C-4 hydroxyl of the galactopyranoside moiety. The location of the sugar is in accord with many biochemical studies.
- Department of Physiology, David Geffen School of Medicine, University of California, Los Angeles, CA 90095, USA.
Organizational Affiliation: