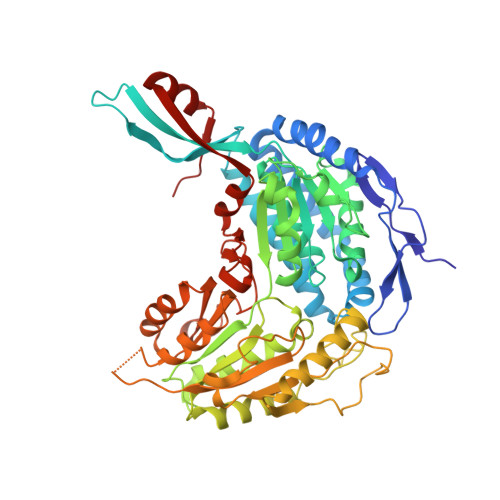
Elucidating the Reaction Mechanism of the Benzoate Oxidation Pathway Encoded Aldehyde Dehydrogenase from Burkholderia Xenovorans Lb400.
Bains, J., Leon, R., Temke, K.G., Boulanger, M.J.(2011) Protein Sci 20: 1048
- PubMed: 21495107 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.639
- Primary Citation Related Structures:
2Y51, 2Y52, 2Y53, 2Y5D - PubMed Abstract:
Oxidation of cis-3,4-dehydroadipyl-CoA semialdehyde to cis-3,4-dehydroadipyl-CoA by the aldehyde dehydrogenase, ALDH(C) (EC.1.2.1.77), is an essential step in the metabolism of benzoate in Burkholderia xenovorans LB400. In a previous study, we established a structural blueprint for this novel group of ALDH enzymes. Here, we build significantly on this initial work and propose a detailed reaction mechanism for ALDH(C) based on comprehensive structural and functional investigations of active site residues. Kinetic analyses reveal essential roles for C296 as the nucleophile and E257 as the associated general base. Structural analyses of E257Q and C296A variants suggest a dynamic charge repulsion relationship between E257 and C296 that contributes to the inherent flexibility of E257 in the native enzyme, which is further regulated by E496 and E167. A proton relay network anchored by E496 and supported by E167 and K168 serves to reset E257 for the second catalytic step. We also propose that E167, which is unique to ALDH(C) and its homologs, serves a critical role in presenting the catalytic water to the newly reset E257 such that the enzyme can proceed with deacylation and product release. Collectively, the reaction mechanism proposed for ALDH(C) promotes a greater understanding of these novel ALDH enzymes, the ALDH super-family in general, and benzoate degradation in B. xenovorans LB400.
- Department of Biochemistry and Microbiology, University of Victoria, Victoria, British Columbia V8W3P6, Canada.
Organizational Affiliation: