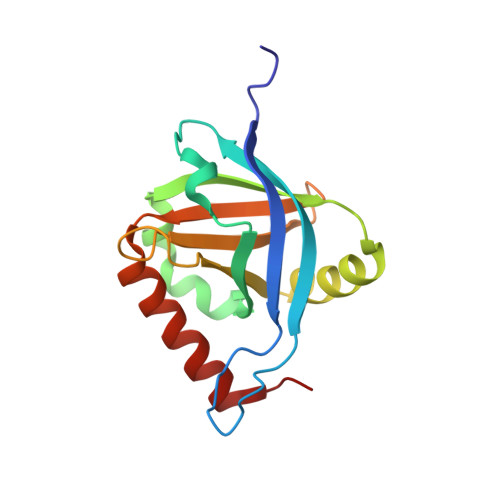
Structure and Vp16 Binding of the Mediator Med25 Activator Interaction Domain.
Vojnic, E., Mourao, A., Seizl, M., Simon, B., Wenzeck, L., Lariviere, L., Baumli, S., Baumgart, K., Meisterernst, M., Sattler, M., Cramer, P.(2011) Nat Struct Mol Biol 18: 404
- PubMed: 21378965 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb.1997
- Primary Citation Related Structures:
2XNF - PubMed Abstract:
Eukaryotic transcription is regulated by interactions between gene-specific activators and the coactivator complex Mediator. Here we report the NMR structure of the Mediator subunit Med25 (also called Arc92) activator interaction domain (ACID) and analyze the structural and functional interaction of ACID with the archetypical acidic transcription activator VP16. Unlike other known activator targets, ACID forms a seven-stranded β-barrel framed by three helices. The VP16 subdomains H1 and H2 bind to opposite faces of ACID and cooperate during promoter-dependent activated transcription in a in vitro system. The activator-binding ACID faces are functionally required and conserved among higher eukaryotes. Comparison with published activator structures reveals that the VP16 activation domain uses distinct interaction modes to adapt to unrelated target surfaces and folds that evolved for activator binding.
- Gene Center and Department of Biochemistry, Center for Integrated Protein Science Munich (CIPSM), Ludwig-Maximilians-Universität München, Munich, Germany.
Organizational Affiliation: