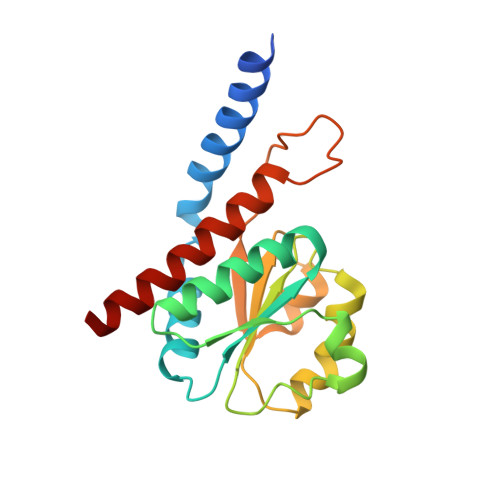
Probing the Active Site of the Sugar Isomerase Domain from E. Coli Arabinose-5-Phosphate Isomerase Via X-Ray Crystallography.
Gourlay, L.J., Sommaruga, S., Nardini, M., Sperandeo, P., Deho, G., Polissi, A., Bolognesi, M.(2010) Protein Sci 19: 2430
- PubMed: 20954237 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.525
- Primary Citation Related Structures:
2XHZ - PubMed Abstract:
Lipopolysaccharide (LPS) biosynthesis represents an underexploited target pathway for novel antimicrobial development to combat the emergence of multidrug-resistant bacteria. A key player in LPS synthesis is the enzyme D-arabinose-5-phosphate isomerase (API), which catalyzes the reversible isomerization of D-ribulose-5-phosphate to D-arabinose-5-phosphate, a precursor of 3-deoxy-D-manno-octulosonate that is an essential residue of the LPS inner core. API is composed of two main domains: an N-terminal sugar isomerase domain (SIS) and a pair of cystathionine-β-synthase domains of unknown function. As the three-dimensional structure of an enzyme is a prerequisite for the rational development of novel inhibitors, we present here the crystal structure of the SIS domain of a catalytic mutant (K59A) of E. coli D-arabinose-5-phosphate isomerase at 2.6-Å resolution. Our structural analyses and comparisons made with other SIS domains highlight several potentially important active site residues. In particular, the crystal structure allowed us to identify a previously unpredicted His residue (H88) located at the mouth of the active site cavity as a possible catalytic residue. On the basis of such structural data, subsequently supported by biochemical and mutational experiments, we confirm the catalytic role of H88, which appears to be a generally conserved residue among two-domain isomerases.
- Dipartimento di Scienze Biomolecolari e Biotecnologie, and CIMAINA, Università di Milano, Via Celoria 26, Milano 20133, Italy.
Organizational Affiliation: