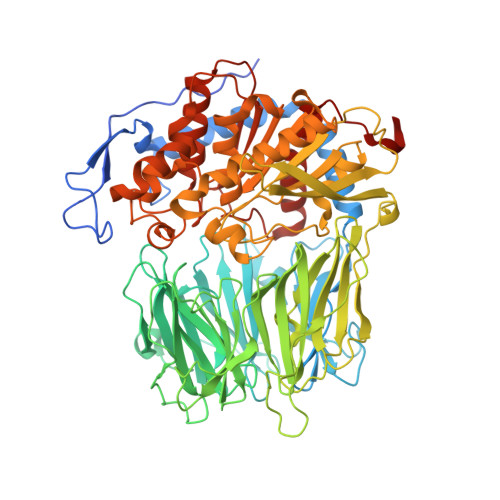
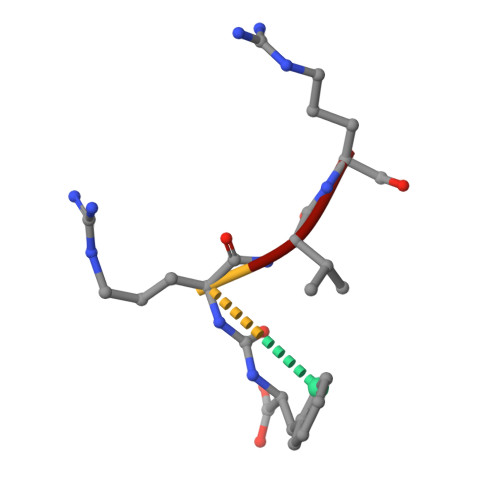
Crystal Structure of Leishmania Major Oligopeptidase B Gives Insight Into the Enzymatic Properties of a Trypanosomatid Virulence Factor.
Mcluskey, K., Paterson, N.G., Bland, N.D., Isaacs, N.W., Mottram, J.C.(2010) J Biological Chem 285: 39249
- PubMed: 20926390 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M110.156679
- Primary Citation Related Structures:
2XE4 - PubMed Abstract:
Oligopeptidase B (OPB) is a serine peptidase with dibasic substrate specificity. It is found in bacteria, plants, and trypanosomatid pathogens, where it has been identified as a virulence factor and potential drug target. In this study we expressed active recombinant Leishmania major OPB and provide the first structure of an oligopeptidase B at high resolution. The crystallographic study reveals that OPB comprises two domains, a catalytic and a propeller domain, linked together by a hinge region. The structure has been determined in complex with the oligopeptide, protease-inhibitor antipain, giving detailed information on the enzyme active site and extended substrate binding pockets. It shows that Glu-621 plays a critical role in the S1 binding pocket and, along with Phe-603, is largely responsible for the enzyme substrate specificity in P1. In the S2 binding pocket, Tyr-499 was shown to be important for substrate stability. The structure also allowed an investigation into the function of residues highlighted in other studies including Glu-623, which was predicted to be involved in the S1 binding pocket but is found forming an inter-domain hydrogen bond. Additional important salt bridges/hydrogen bonds between the two domains were observed, highlighting the significance of the domain interface in OPB. This work provides a foundation for the study of the role of OPBs as virulence factors in trypanosomatids. It could facilitate the development of specific OPB inhibitors with therapeutic potential by exploiting its unique substrate recognition properties as well as providing a model for OPBs in general.
- Westchem School of Chemistry, University of Glasgow, Glasgow G12 8TA, Scotland, United Kingdom. Karen.McLuskey@glasgow.ac.uk
Organizational Affiliation: