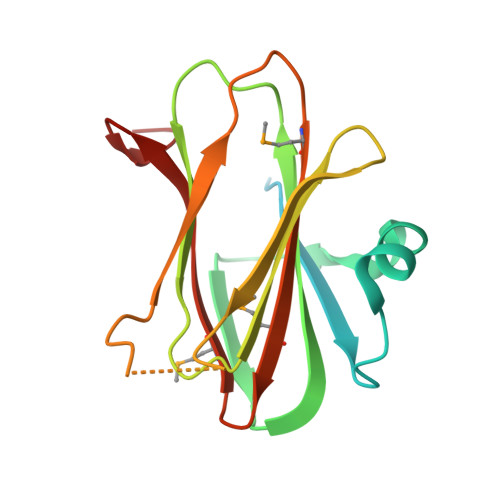
Distinct Oligomeric Forms of the Pseudomonas Aeruginosa Rets Sensor Domain Modulate Accessibility to the Ligand Binding Site.
Vincent, F., Round, A., Reynaud, A., Bordi, C., Filloux, A., Bourne, Y.(2010) Environ Microbiol 12: 1775
- PubMed: 20553556 Search on PubMed
- DOI: https://doi.org/10.1111/j.1462-2920.2010.02264.x
- Primary Citation Related Structures:
2XBZ - PubMed Abstract:
Bacterial two-component regulatory systems (TCSs) sense environmental stimuli to adapt the lifestyle of microbial populations. For many TCSs the stimulus is a ligand of unknown chemical nature. Pseudomonas aeruginosa utilizes the closely related RetS and LadS sensor kinases to switch between acute and chronic infections. These sensor proteins antagonistically mediate biofilm formation through communication with a central TCS, GacA/GacS. Recently, it was shown that RetS modulates the GacS sensor activity by forming RetS/GacS heterodimers. LadS and RetS are hybrid sensors with a signalling domain consisting of a 7-transmembrane (7TMR) region and a periplasmic sensor domain (diverse intracellular signalling module extracellular 2, DISMED2). The 2.65 A resolution crystal structure of RetS DISMED2, called RetSp, reveals three distinct oligomeric states capable of domain swapping. The RetSp structure also displays two putative ligand binding sites. One is equivalent to the analogous site in the structurally-related carbohydrate binding module (CBM) but the second site is located at a dimer interface. These observations highlight the modular architecture and assembly of the RetSp fold and give clues on how homodimerization of RetS could be modulated upon ligand binding to control formation of a RetS/GacS heterodimer. Modelling the DISMED2 of LadS reveals conservation of only one ligand binding site, suggesting a distinct mechanism underlying the activity of this sensor kinase.
- Architecture et Fonction des Macromolécules Biologiques, UMR6098, CNRS et Universités Aix-Marseille I et II, 163 avenue de Luminy, 13288 Marseille, France. florence.vincent@afmb.univ-mrs.fr
Organizational Affiliation: