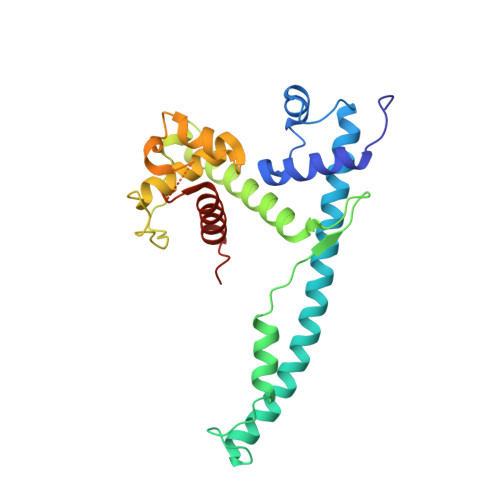
The DNA-Binding Domain of the Chd1 Chromatin- Remodelling Enzyme Contains Sant and Slide Domains.
Ryan, D.P., Sundaramoorthy, R., Martin, D., Singh, V., Owen-Hughes, T.(2011) EMBO J 30: 2596
- PubMed: 21623345 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/emboj.2011.166
- Primary Citation Related Structures:
2XB0 - PubMed Abstract:
The ATP-dependent chromatin-remodelling enzyme Chd1 is a 168-kDa protein consisting of a double chromodomain, Snf2-related ATPase domain, and a C-terminal DNA-binding domain. Here, we show the DNA-binding domain is required for Saccharomyces cerevisiae Chd1 to bind and remodel nucleosomes. The crystal structure of this domain reveals the presence of structural homology to SANT and SLIDE domains previously identified in ISWI remodelling enzymes. The presence of these domains in ISWI and Chd1 chromatin-remodelling enzymes may provide a means of efficiently harnessing the action of the Snf2-related ATPase domain for the purpose of nucleosome spacing and provide an explanation for partial redundancy between these proteins. Site directed mutagenesis was used to identify residues important for DNA binding and generate a model describing the interaction of this domain with DNA. Through inclusion of Chd1 sequences in homology searches SLIDE domains were identified in CHD6-9 proteins. Point mutations to conserved amino acids within the human CHD7 SLIDE domain have been identified in patients with CHARGE syndrome.
- Wellcome Trust Centre for Gene Regulation and Expression, College of Life Sciences, University of Dundee, Dundee, UK.
Organizational Affiliation: