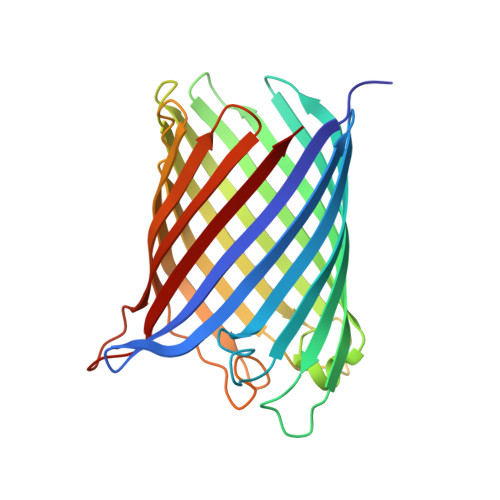
Correlation between the Ompg Secondary Structure and its Ph-Dependent Alterations Monitored by Ftir.
Korkmaz-Ozkan, F., Koster, S., Kuhlbrandt, W., Mantele, W., Yildiz, O.(2010) J Mol Biology 401: 56
- PubMed: 20561532 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2010.06.015
- Primary Citation Related Structures:
2X9K - PubMed Abstract:
The channel activity of the outer-membrane protein G (OmpG) from Escherichia coli is pH-dependent. To investigate the role of the histidine pair His231/His261 in triggering channel opening and closing, we mutated both histidines to alanines and cysteines. Fourier transform infrared spectra revealed that the OmpG mutants stay-independent of pH-in an open conformation. Temperature ramp experiments indicate that the mutants are as stable as the open state of wild-type OmpG. The X-ray structure of the alanine-substituted OmpG mutant obtained at pH 6.5 confirms the constitutively open conformation. Compared to previous structures of the wild-type protein in the open and closed conformation, the mutant structure shows a difference in the extracellular loop L6 connecting beta-strands S12 and S13. A deletion of amino acids 220-228, which are thought to block the channel at low pH in wild-type OmpG, indicates conformational changes, which might be triggered by His231/His261.
- Institute of Biophysics, Goethe-University, Max-von-Laue-Str. 1, D-60438 Frankfurt am Main, Germany.
Organizational Affiliation: