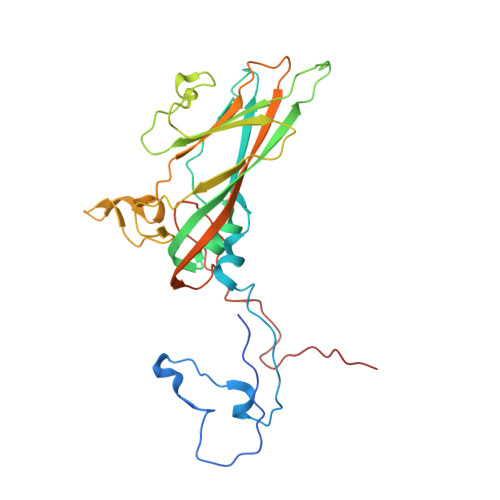
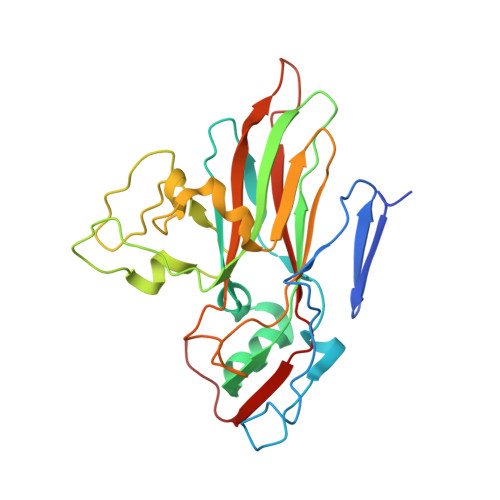
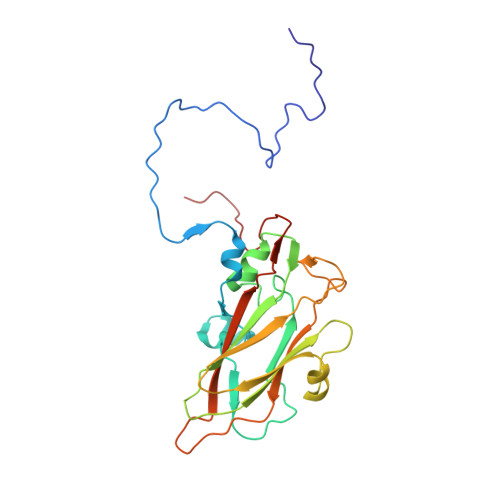
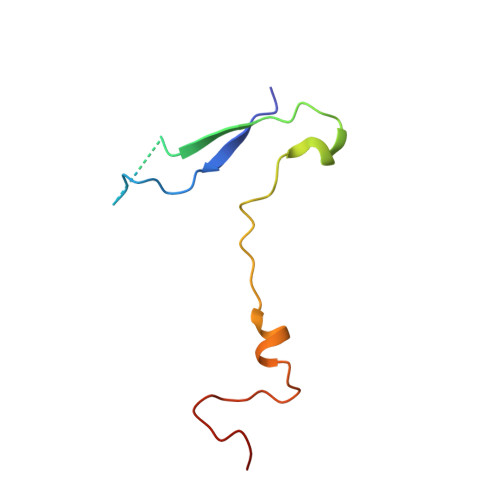
Interaction of Decay-Accelerating Factor with Echovirus 7.
Plevka, P., Hafenstein, S., Harris, K.G., Cifuente, J.O., Zhang, Y., Bowman, V.D., Chipman, P.R., Bator, C.M., Lin, F., Medof, M.E., Rossmann, M.G.(2010) J Virol 84: 12665
- PubMed: 20881044
- DOI: https://doi.org/10.1128/JVI.00837-10
- Primary Citation of Related Structures:
2X5I, 3IYP - PubMed Abstract:
Echovirus 7 (EV7) belongs to the Enterovirus genus within the family Picornaviridae. Many picornaviruses use IgG-like receptors that bind in the viral canyon and are required to initiate viral uncoating during infection. However, in addition, some of the enteroviruses use an alternative or additional receptor that binds outside the canyon. Decay-accelerating factor (DAF) has been identified as a cellular receptor for EV7. The crystal structure of EV7 has been determined to 3.1-Å resolution and used to interpret the 7.2-Å-resolution cryo-electron microscopy reconstruction of EV7 complexed with DAF. Each DAF binding site on EV7 is near a 2-fold icosahedral symmetry axis, which differs from the binding site of DAF on the surface of coxsackievirus B3, indicating that there are independent evolutionary processes by which DAF was selected as a picornavirus accessory receptor. This suggests that there is an advantage for these viruses to recognize DAF during the initial process of infection.
- Department of Biological Sciences, Purdue University, 915 W. State Street, West Lafayette, IN 47907-2054, USA.
Organizational Affiliation: