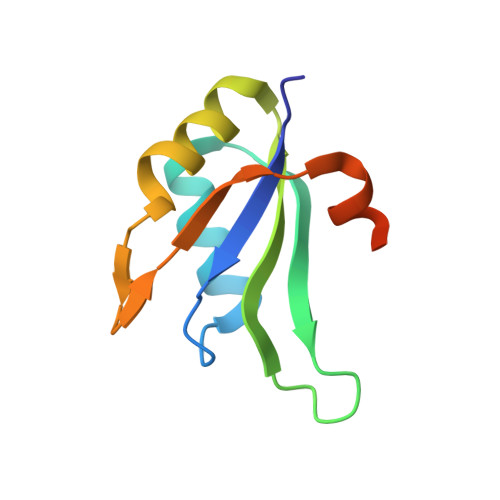
Structure of the RNA15 Rrm-RNA Complex Reveals the Molecular Basis of Gu Specificity in Transcriptional 3-End Processing Factors.
Pancevac, C., Goldstone, D.C., Ramos, A., Taylor, I.A.(2010) Nucleic Acids Res 38: 3119
- PubMed: 20097654
- DOI: https://doi.org/10.1093/nar/gkq002
- Primary Citation Related Structures:
2X1A, 2X1B, 2X1F - PubMed Abstract:
Rna15 is a core subunit of cleavage factor IA (CFIA), an essential transcriptional 3'-end processing factor from Saccharomyces cerevisiae. CFIA is required for polyA site selection/cleavage targeting RNA sequences that surround polyadenylation sites in the 3'-UTR of RNA polymerase-II transcripts. RNA recognition by CFIA is mediated by an RNA recognition motif (RRM) contained in the Rna15 subunit of the complex. We show here that Rna15 has a strong and unexpected preference for GU containing RNAs and reveal the molecular basis for a base selectivity mechanism that accommodates G or U but discriminates against C and A bases. This mode of base selectivity is rather different to that observed in other RRM-RNA structures and is structurally conserved in CstF64, the mammalian counterpart of Rna15. Our observations provide evidence for a highly conserved mechanism of base recognition amongst the 3'-end processing complexes that interact with the U-rich or U/G-rich elements at 3'-end cleavage/polyadenylation sites.
- Division of Molecular Structure, MRC National Institute for Medical Research, The Ridgeway, Mill Hill, London NW7 1AA, UK.
Organizational Affiliation: