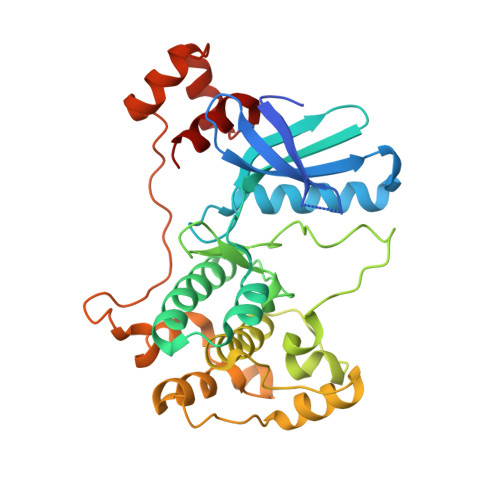
Structure and Function of Polarity-Inducing Kinase Family Mark/Par-1 within the Branch of Ampk/Snf1-Related Kinases.
Marx, A., Nugoor, C., Panneerselvam, S., Mandelkow, E.(2010) FASEB J 24: 1637
- PubMed: 20071654 Search on PubMed
- DOI: https://doi.org/10.1096/fj.09-148064
- Primary Citation Related Structures:
2WZJ - PubMed Abstract:
Kinases of the MARK/Par-1 family of S/T protein kinases are regulators of diverse cellular processes in Caenorhabditis elegans, Drosophila, yeast, and mammalian cells. They are involved in nematode embryogenesis, epithelial cell polarization, cell signaling, and neuronal differentiation. MARK phosphorylates microtubule-associated proteins such as tau and is a key regulator of microtubule-based intracellular transport. Hyperphosphorylation of tau causes defects in neuronal transport and may induce abnormal aggregation of tau in Alzheimer disease and other tauopathies. Recent high-resolution structure analysis of MARK fragments covering the kinase domain and accessory regulatory domains has revealed important details regarding the autoregulation of MARK, but their interpretation has remained controversial. Here we focus on the structural aspects of MARK activity and autoregulation. Comparison of the available MARK structures with related kinases of the AMPK family and with new structures of MARK isoforms (MARK2 and 3) reveals unexpected structural similarities between these kinases that may help to resolve the existing controversies.
- Max Planck Unit for Structural Molecular Biology, Hamburg, Germany. marx@mpasmb.desy.de
Organizational Affiliation: