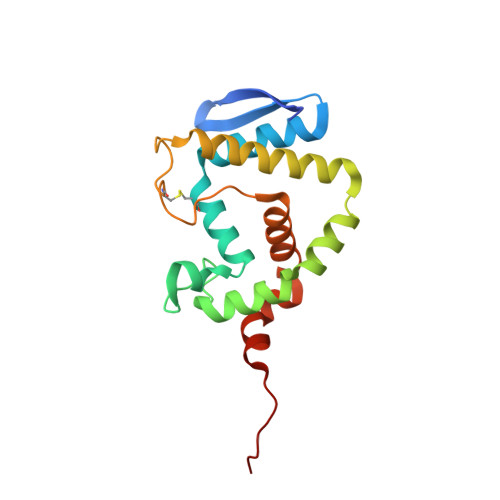
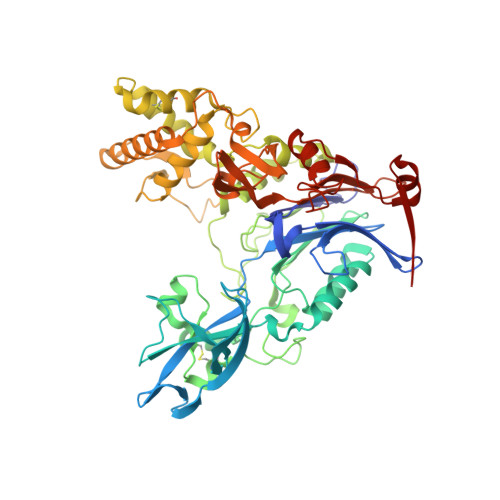
The Quorum-Quenching N-Acyl Homoserine Lactone Acylase Pvdq is an Ntn-Hydrolase with an Unusual Substrate-Binding Pocket
Bokhove, M., Nadal Jimenez, P., Quax, W.J., Dijkstra, B.W.(2010) Proc Natl Acad Sci U S A 107: 686
- PubMed: 20080736 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0911839107
- Primary Citation Related Structures:
2WYB, 2WYC, 2WYD, 2WYE - PubMed Abstract:
In many Gram-negative pathogens, their virulent behavior is regulated by quorum sensing, in which diffusible signals such as N-acyl homoserine lactones (AHLs) act as chemical messaging compounds. Enzymatic degradation of these diffusible signals by, e.g., lactonases or amidohydrolases abolishes AHL regulated virulence, a process known as quorum quenching. Here we report the first crystal structure of an AHL amidohydrolase, the AHL acylase PvdQ from Pseudomonas aeruginosa. PvdQ has a typical alpha/beta heterodimeric Ntn-hydrolase fold, similar to penicillin G acylase and cephalosporin acylase. However, it has a distinct, unusually large, hydrophobic binding pocket, ideally suited to recognize C12 fatty acid-like chains of AHLs. Binding of a C12 fatty acid or a 3-oxo-C12 fatty acid induces subtle conformational changes to accommodate the aliphatic chain. Furthermore, the structure of a covalent ester intermediate identifies Serbeta1 as the nucleophile and Asnbeta269 and Valbeta70 as the oxyanion hole residues in the AHL degradation process. Our structures show the versatility of the Ntn-hydrolase scaffold and can serve as a structural paradigm for Ntn-hydrolases with similar substrate preference. Finally, the quorum-quenching capabilities of PvdQ may be utilized to suppress the quorum-sensing machinery of pathogens.
- Laboratory of Biophysical Chemistry, University of Groningen, Nijenborgh 4, 9747 AG Groningen, The Netherlands.
Organizational Affiliation: