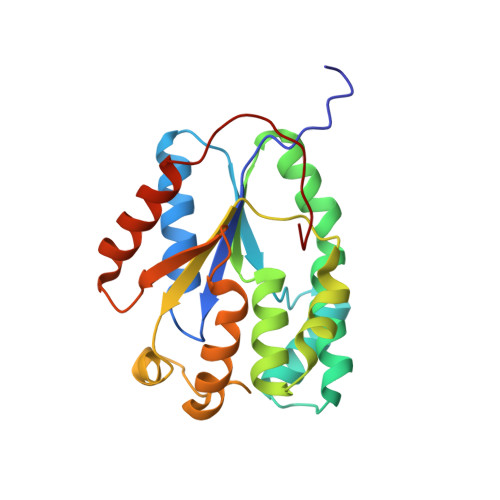
Structural Basis for the Efficient Phosphorylation of Aztmp and Dgmp by Plasmodium Falciparum Type I Thymidylate Kinase.
Whittingham, J.L., Carrero-Lerida, J., Brannigan, J.A., Ruiz-Perez, L.M., Silva, A.P.G., Fogg, M.J., Wilkinson, A.J., Gilbert, I.H., Wilson, K.S., Gonzalez-Pacanowska, D.(2010) Biochem J 428: 499
- PubMed: 20353400 Search on PubMed
- DOI: https://doi.org/10.1042/BJ20091880
- Primary Citation Related Structures:
2WWF, 2WWG, 2WWH, 2WWI - PubMed Abstract:
Plasmodium falciparum is the causative agent of malaria, a disease where new drug targets are required due to increasing resistance to current anti-malarials. TMPK (thymidylate kinase) is a good candidate as it is essential for the synthesis of dTTP, a critical precursor of DNA and has been much studied due to its role in prodrug activation and as a drug target. Type I TMPKs, such as the human enzyme, phosphorylate the substrate AZT (3'-azido-3'-deoxythymidine)-MP (monophosphate) inefficiently compared with type II TMPKs (e.g. Escherichia coli TMPK). In the present paper we report that eukaryotic PfTMPK (P. falciparum TMPK) presents sequence features of a type I enzyme yet the kinetic parameters for AZT-MP phosphorylation are similar to those of the highly efficient E. coli enzyme. Structural information shows that this is explained by a different juxtaposition of the P-loop and the azide of AZT-MP. Subsequent formation of the transition state requires no further movement of the PfTMPK P-loop, with no steric conflicts for the azide moiety, allowing efficient phosphate transfer. Likewise, we present results that confirm the ability of the enzyme to uniquely accept dGMP as a substrate and shed light on the basis for its wider substrate specificity. Information resulting from two ternary complexes (dTMP-ADP and AZT-MP-ADP) and a binary complex with the transition state analogue AP5dT [P1-(5'-adenosyl)-P5-(5'-thymidyl) pentaphosphate] all reveal significant differences with the human enzyme, notably in the lid region and in the P-loop which may be exploited in the rational design of Plasmodium-specific TMPK inhibitors with therapeutic potential.
- Structural Biology Laboratory, Department of Chemistry, University of York, Heslington, York YO105YW, UK.
Organizational Affiliation: