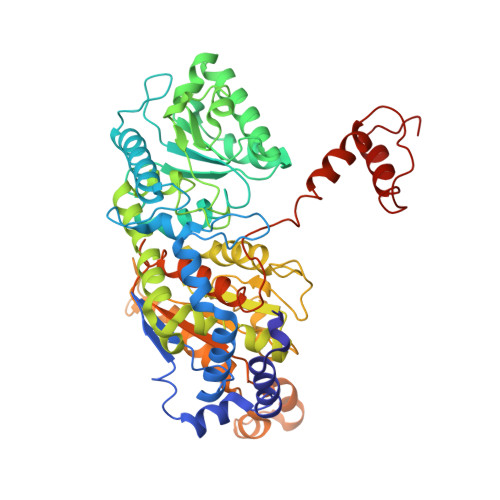
Structural Studies of Phosphoglucose Isomerase from Mycobacterium Tuberculosis H37Rv
Anand, K., Mathur, D., Anant, A., Garg, L.C.(2010) Acta Crystallogr Sect F Struct Biol Cryst Commun 66: 490
- PubMed: 20445242 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309110011656
- Primary Citation Related Structures:
2WU8 - PubMed Abstract:
Phosphoglucose isomerase (PGI) plays a key role in both glycolysis and gluconeogenesis inside the cell, whereas outside the cell it exhibits cytokine properties. PGI is also known to act as an autocrine motility factor, a neuroleukin agent and a differentiation and maturation mediator. Here, the first crystal structure of PGI from Mycobacterium tuberculosis H37Rv (Mtb) is reported. The structure was refined at 2.25 A resolution and revealed the presence of one molecule in the asymmetric unit with two globular domains. As known previously, the active site of Mtb PGI contains conserved residues including Glu356, Glu216 and His387 (where His387 is from the neighbouring molecule). The crystal structure of Mtb PGI was observed to be rather more similar to human PGI than other nonbacterial PGIs, with only a few differences being detected in the loops, arm and hook regions of the human and Mtb PGIs, suggesting that the M. tuberculosis enzyme uses the same enzyme mechanism.
- European Molecular Biology Laboratory Heidelberg, Structural and Computational Biology Unit, Meyerhof Strasse 1, D-69117 Heidelberg, Germany. anand@embl.de,lalit@nii.res.in
Organizational Affiliation: