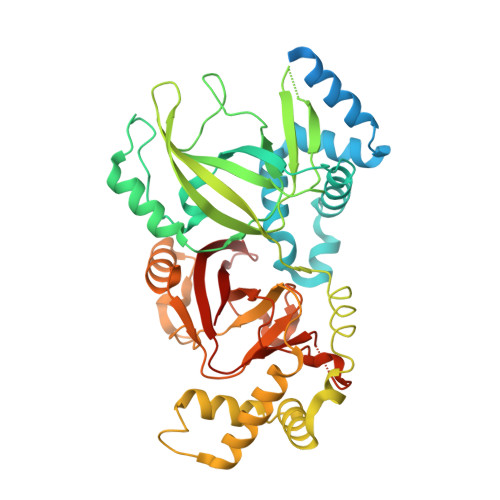
Structural Basis for Substrate Recognition in the Enzymatic Component of Adp-Ribosyltransferase Toxin Cdta from Clostridium Difficile.
Sundriyal, A., Roberts, A.K., Shone, C.C., Acharya, K.R.(2009) J Biological Chem 284: 28713
- PubMed: 19692332 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M109.043018
- Primary Citation Related Structures:
2WN4, 2WN5, 2WN6, 2WN7, 2WN8 - PubMed Abstract:
ADP-ribosylation is one of the favored modes of cell intoxication employed by several bacteria. Clostridium difficile is recognized to be an important nosocomial pathogen associated with considerable morbidity and attributable mortality. Along with its two well known toxins, Toxin A and Toxin B, it produces an ADP-ribosylating toxin that targets monomeric actin of the target cell. Like other Clostridial actin ADP-ribosylating toxins, this binary toxin, known as C. difficile toxin (CDT), is composed of two subunits, CDTa and CDTb. In this study, we present high resolution crystal structures of CDTa in its native form (at pH 4.0, 8.5, and 9.0) and in complex with ADP-ribose donors, NAD and NADPH (at pH 9.0). The crystal structures of the native protein show "pronounced conformational flexibility" confined to the active site region of the protein and "enhanced" disorder at low pH, whereas the complex structures highlight significant differences in "ligand specificity" compared with the enzymatic subunit of a close homologue, Clostridium perfringens iota toxin. Specifically in CDTa, two of the suggested catalytically important residues (Glu-385 and Glu-387) seem to play no role or a less important role in ligand binding. These structural data provide the first detailed information on protein-donor substrate complex stabilization in CDTa, which may have implications in understanding CDT recognition.
- Department of Biology and Biochemistry, University of Bath, Claverton Down, Bath BA2 7AY, United Kingdom.
Organizational Affiliation: