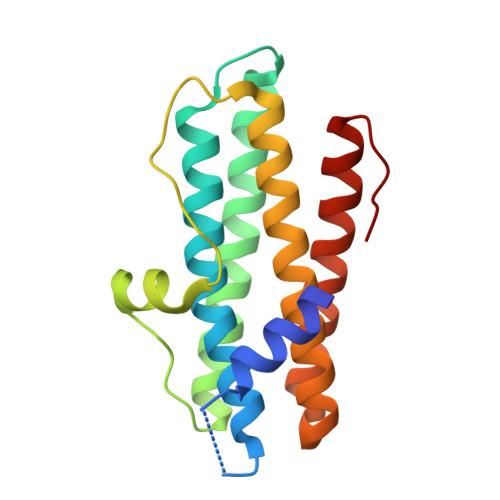
Crystal Structures of Streptococcus Pyogenes Dpr Reveal a Dodecameric Iron-Binding Protein with a Ferroxidase Site.
Haikarainen, T., Tsou, C.-C., Wu, J.-J., Papageorgiou, A.C.(2010) J Biol Inorg Chem 15: 183
- PubMed: 19727858 Search on PubMed
- DOI: https://doi.org/10.1007/s00775-009-0582-9
- Primary Citation Related Structures:
2WLA, 2WLU - PubMed Abstract:
DNA-binding protein from starved cells (Dps)-like proteins are key factors involved in oxidative stress protection in bacteria. They bind and oxidize iron, thus preventing the formation of harmful reactive oxygen species that can damage biomolecules, particularly DNA. Dps-like proteins are composed of 12 identical subunits assembled in a spherical structure with a hollow central cavity. The iron oxidation occurs at 12 intersubunit sites located at dimer interfaces. Streptococcus pyogenes Dps-like peroxide resistance protein (Dpr) has been previously found to protect the catalase-lacking S. pyogenes bacterium from oxidative stress. We have determined the crystal structure of S. pyogenes Dpr, the second Dpr structure from a streptococcal bacterium, in iron-free and iron-bound forms at 2.0- and 1.93-A resolution, respectively. The iron binds to well-conserved sites at dimer interfaces and is coordinated directly to Asp77 and Glu81 from one monomer, His50 from a twofold symmetry-related monomer, a glycerol molecule, and a water molecule. Upon iron binding, Asp77 and Glu81 change conformation. Site-directed mutagenesis of active-site residues His50, His62, Asp66, Asp77, and Glu81 to Ala revealed a dramatic decrease in iron incorporation. A short helix at the N-terminal was found in a different position compared with other Dps-like proteins. Two types of pores were identified in the dodecamer. Although the N-terminal pore was found to be similar to that of other Dps-like proteins, the C-terminal pore was found to be blocked by bulky Tyr residues instead of small residues present in other Dps-like proteins.
- Turku Centre for Biotechnology, University of Turku and Abo Akademi University, Turku, Finland.
Organizational Affiliation: