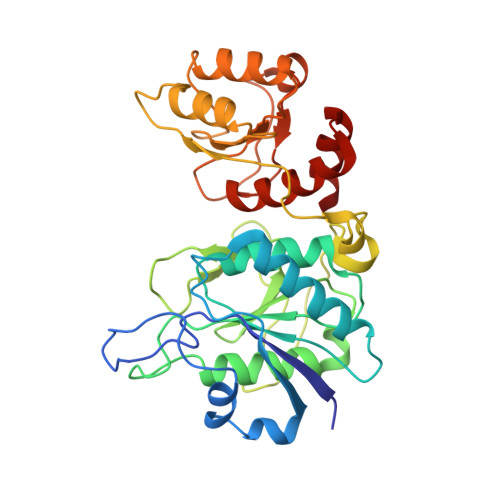
Structure of Helicobacter Pylori L-Asparaginase at 1.4 A Resolution
Dhavala, P., Papageorgiou, A.C.(2009) Acta Crystallogr D Biol Crystallogr 65: 1253
- PubMed: 19966411 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444909038244
- Primary Citation Related Structures:
2WLT, 2WT4 - PubMed Abstract:
Bacterial L-asparaginases have been used in the treatment of childhood acute lymphoblastic leukaemia for over 30 years. Their therapeutic effect is based on their ability to catalyze the conversion of L-asparagine, an essential amino acid in certain tumours, to L-aspartic acid and ammonia. Two L-asparaginases, one from Escherichia coli and the other from Erwinia chrysanthemi, have been widely employed in clinical practice as anti-leukaemia drugs. However, L-asparaginases are also able to cause severe side effects owing to their intrinsic glutaminase activity. Helicobacter pylori L-asparaginase (HpA) has been reported to have negligible glutaminase activity. To gain insight into the properties of HpA, its crystal structure in the presence of L-aspartate was determined to 1.4 A resolution, which is one of the highest resolutions obtained for an L-asparaginase structure. The final structure has an R(cryst) of 12.6% (R(free) = 16.9%) with good stereochemistry. A detailed analysis of the active site showed major differences in the active-site flexible loop and in the 286-297 loop from the second subunit, which is involved in active-site formation. Accordingly, Glu289, Asn255 and Gln63 are suggested to play roles in modulating the accessibility of the active site. Overall, the structural comparison revealed that HpA has greater structural similarity to E. coli L-asparaginase than to any other L-asparaginase, including Er. carotovora L-asparaginase, despite the fact that the latter is also characterized by low glutaminase activity.
- Turku Centre for Biotechnology, University of Turku and Abo Akademi University, Finland.
Organizational Affiliation: