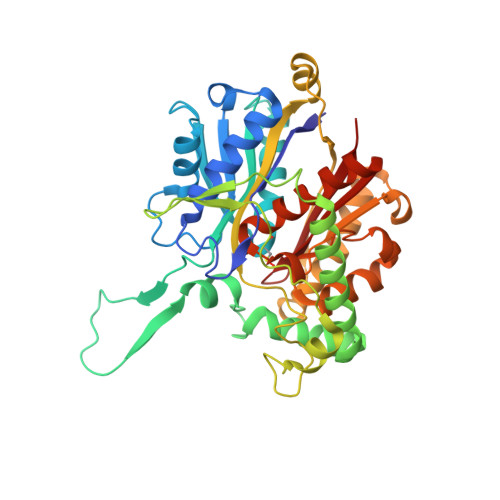
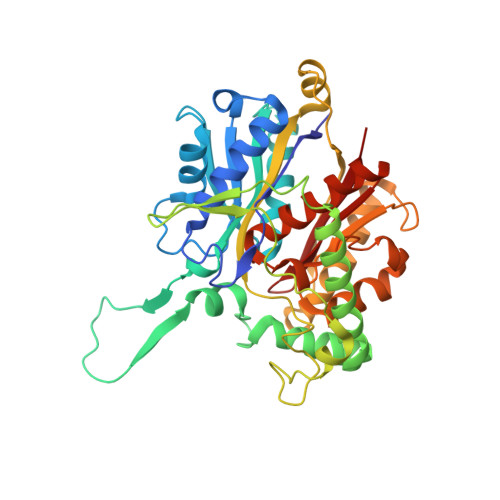
The Thiolase Reaction Mechanism: The Importance of Asn316 and His348 for Stabilizing the Enolate Intermediate of the Claisen Condensation.
Merilainen, G., Poikela, V., Kursula, P., Wierenga, R.K.(2009) Biochemistry 48: 11011
- PubMed: 19842716 Search on PubMed
- DOI: https://doi.org/10.1021/bi901069h
- Primary Citation Related Structures:
2WKT, 2WKU, 2WKV, 2WL4, 2WL5, 2WL6 - PubMed Abstract:
The biosynthetic thiolase catalyzes a Claisen condensation reaction between acetyl-CoA and the enzyme acetylated at Cys89. Two oxyanion holes facilitate this catalysis: oxyanion hole I stabilizes the enolate intermediate generated from acetyl-CoA, whereas oxyanion hole II stabilizes the tetrahedral intermediate of the acetylated enzyme. The latter intermediate is formed when the alpha-carbanion of acetyl-CoA enolate reacts with the carbonyl carbon of acetyl-Cys89, after which C-C bond formation is completed. Oxyanion hole II is made of two main chain peptide NH groups, whereas oxyanion hole I is formed by a water molecule (Wat82) and NE2(His348). Wat82 is anchored in the active site by an optimal set of hydrogen bonding interactions, including a hydrogen bond to ND2(Asn316). Here, the importance of Asn316 and His348 for catalysis has been studied; in particular, the properties of the N316D, N316A, N316H, H348A, and H348N variants have been determined. For the N316D variant, no activity could be detected. For each of the remaining variants, the k(cat)/K(m) value for the Claisen condensation catalysis is reduced by a factor of several hundred, whereas the thiolytic degradation catalysis is much less affected. The crystal structures of the variants show that the structural changes in the active site are minimal. Our studies confirm that oxyanion hole I is critically important for the condensation catalysis. Removing either one of the hydrogen bond donors causes the loss of at least 3.4 kcal/mol of transition state stabilization. It appears that in the thiolytic degradation direction, oxyanion hole I is not involved in stabilizing the transition state of its rate limiting step. However, His348 has a dual role in the catalytic cycle, contributing to oxyanion hole I and activating Cys89. The analysis of the hydrogen bonding interactions in the very polar catalytic cavity shows the importance of two conserved water molecules, Wat82 and Wat49, for the formation of oxyanion hole I and for influencing the reactivity of the catalytic base, Cys378, respectively. Cys89, Asn316, and His348 form the CNH-catalytic triad of the thiolase superfamily. Our findings are also discussed in the context of the importance of this triad for the catalytic mechanism of other enzymes of the thiolase superfamily.
- Department of Biochemistry, University of Oulu, P.O. Box 3000, 90014 Oulu, Finland.
Organizational Affiliation: