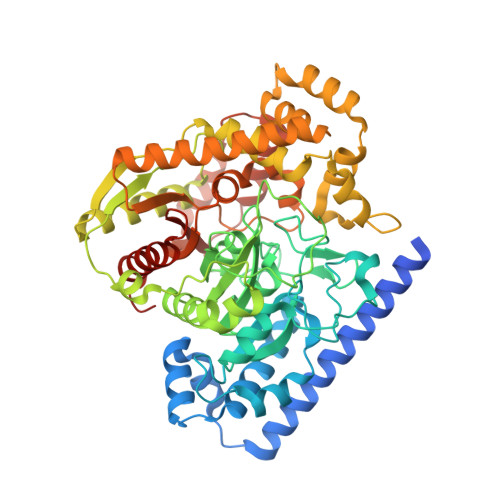
Binding and Inactivation Mechanism of a Humanized Fatty Acid Amide Hydrolase by Alpha-Ketoheterocycle Inhibitors Revealed from Cocrystal Structures.
Mileni, M., Garfunkle, J., Demartino, J.K., Cravatt, B.F., Boger, D.L., Stevens, R.C.(2009) J Am Chem Soc 131: 10497
- PubMed: 19722626 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/ja902694n
- Primary Citation Related Structures:
2WJ1, 2WJ2 - PubMed Abstract:
The cocrystal X-ray structures of two isomeric alpha-ketooxazole inhibitors (1 (OL-135) and 2) bound to fatty acid amide hydrolase (FAAH), a key enzymatic regulator of endocannabinoid signaling, are disclosed. The active site catalytic Ser241 is covalently bound to the inhibitors' electrophilic carbonyl groups, providing the first structures of FAAH bound to an inhibitor as a deprotonated hemiketal mimicking the enzymatic tetrahedral intermediate. The work also offers a detailed view of the oxyanion hole and an exceptional "in-action" depiction of the unusual Ser-Ser-Lys catalytic triad. These structures capture the first picture of inhibitors that span the active site into the cytosolic port providing new insights that help to explain FAAH's interaction with substrate leaving groups and their role in modulating inhibitor potency and selectivity. The role for the activating central heterocycle is clearly defined and distinguished from that observed in prior applications with serine proteases, reconciling the large electronic effect of attached substituents found unique to this class of inhibitors with FAAH. Additional striking active site flexibility is seen upon binding of the inhibitors, providing insights into the existence of a now well-defined membrane access channel with the disappearance of a spatially independent portion of the acyl chain-binding pocket. Finally, comparison of the structures of OL-135 (1) and its isomer 2 indicates that they bind identically to FAAH, albeit with reversed orientations of the central activating heterocycle, revealing that the terminal 2-pyridyl substituent and the acyl chain phenyl group provide key anchoring interactions and confirming the distinguishing role of the activating oxazole.
- Departments of Molecular Biology, The Scripps Research Institute, 10550 North Torrey Pines Road, La Jolla, California 92037, USA.
Organizational Affiliation: