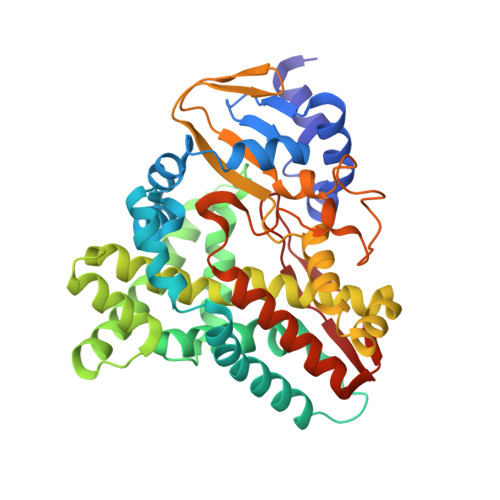
The 1.5-A Structure of Xpla-Heme, an Unusual Cytochrome P450 Heme Domain that Catalyzes Reductive Biotransformation of Royal Demolition Explosive.
Sabbadin, F., Jackson, R., Haider, K., Tampi, G., Turkenburg, J.P., Hart, S., Bruce, N.C., Grogan, G.(2009) J Biological Chem 284: 28467
- PubMed: 19692330 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M109.031559
- Primary Citation Related Structures:
2WIV, 2WIY - PubMed Abstract:
XplA is a cytochrome P450 of unique structural organization, consisting of a heme-domain that is C-terminally fused to its native flavodoxin redox partner. XplA, along with flavodoxin reductase XplB, has been shown to catalyze the breakdown of the nitramine explosive and pollutant hexahydro-1,3,5-trinitro-1,3,5-triazine (royal demolition explosive) by reductive denitration. The structure of the heme domain of XplA (XplA-heme) has been solved in two crystal forms: as a dimer in space group P2(1) to a resolution of 1.9 A and as a monomer in space group P2(1)2(1)2 to a resolution of 1.5 A, with the ligand imidazole bound at the heme iron. Although it shares the overall fold of cytochromes P450 of known structure, XplA-heme is unusual in that the kinked I-helix that traverses the distal face of the heme is broken by Met-394 and Ala-395 in place of the well conserved Asp/Glu plus Thr/Ser, important in oxidative P450s for the scission of the dioxygen bond prior to substrate oxygenation. The heme environment of XplA-heme is hydrophobic, featuring a cluster of three methionines above the heme, including Met-394. Imidazole was observed bound to the heme iron and is in close proximity to the side chain of Gln-438, which is situated over the distal face of the heme. Imidazole is also hydrogen-bonded to a water molecule that sits in place of the threonine side-chain hydroxyl exemplified by Thr-252 in Cyt-P450cam. Both Gln-438 --> Ala and Ala-395 --> Thr mutants of XplA-heme displayed markedly reduced activity compared with the wild type for royal demolition explosive degradation when combined with surrogate electron donors.
- York Structural Biology Laboratory, Department of Biology, University of York, York YO10 5YW, United Kingdom; Centre for Novel Agricultural Products, Department of Biology, University of York, York YO10 5YW, United Kingdom.
Organizational Affiliation: