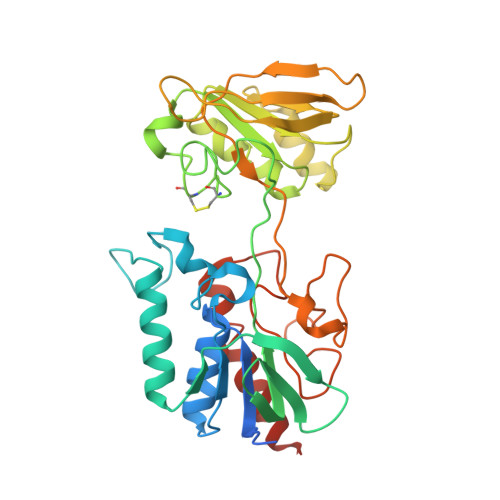
Structure of Hordeum Vulgare Nadph-Dependent Thioredoxin Reductase 2. Unwinding the Reaction Mechanism.
Kirkensgaard, K.G., Hagglund, P., Finnie, C., Svensson, B., Henriksen, A.(2009) Acta Crystallogr D Biol Crystallogr 65: 932
- PubMed: 19690371
- DOI: https://doi.org/10.1107/S0907444909021817
- Primary Citation Related Structures:
2WHD - PubMed Abstract:
Thioredoxins (Trxs) are protein disulfide reductases that regulate the intracellular redox environment and are important for seed germination in plants. Trxs are in turn regulated by NADPH-dependent thioredoxin reductases (NTRs), which provide reducing equivalents to Trx using NADPH to recycle Trxs to the active form. Here, the first crystal structure of a cereal NTR, HvNTR2 from Hordeum vulgare (barley), is presented, which is also the first structure of a monocot plant NTR. The structure was determined at 2.6 A resolution and refined to an R(cryst) of 19.0% and an R(free) of 23.8%. The dimeric protein is structurally similar to the structures of AtNTR-B from Arabidopsis thaliana and other known low-molecular-weight NTRs. However, the relative position of the two NTR cofactor-binding domains, the FAD and the NADPH domains, is not the same. The NADPH domain is rotated by 25 degrees and bent by a 38% closure relative to the FAD domain in comparison with AtNTR-B. The structure may represent an intermediate between the two conformations described previously: the flavin-oxidizing (FO) and the flavin-reducing (FR) conformations. Here, analysis of interdomain contacts as well as phylogenetic studies lead to the proposal of a new reaction scheme in which NTR-Trx interactions mediate the FO to FR transformation.
- Carlsberg Laboratory, Denmark.
Organizational Affiliation: