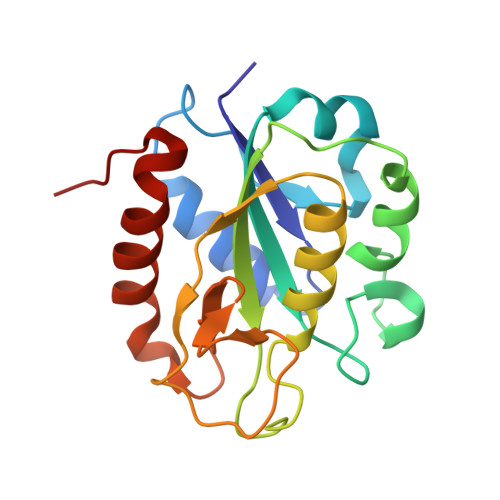
Structural and Phylogenetic Analysis of Rhodobacter Capsulatus Niff: Uncovering General Features of Nitrogen-Fixation (Nif)-Flavodoxins.
Perez-Dorado, I., Bortolotti, A., Cortez, N., Hermoso, J.A.(2013) Int J Mol Sci 14: 1152
- PubMed: 23303276 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/ijms14011152
- Primary Citation Related Structures:
2WC1 - PubMed Abstract:
Analysis of the crystal structure of NifF from Rhodobacter capsulatus and its homologues reported so far reflects the existence of unique structural features in nif flavodoxins: a leucine at the re face of the isoalloxazine, an eight-residue insertion at the C-terminus of the 50's loop and a remarkable difference in the electrostatic potential surface with respect to non-nif flavodoxins. A phylogenetic study on 64 sequences from 52 bacterial species revealed four clusters, including different functional prototypes, correlating the previously defined as "short-chain" with the firmicutes flavodoxins and the "long-chain" with gram-negative species. The comparison of Rhodobacter NifF structure with other bacterial flavodoxin prototypes discloses the concurrence of specific features of these functional electron donors to nitrogenase.
- Department of Crystallography and Structural Biology, Institute of Physical-Chemistry "Rocasolano", CSIC, Serrano 119, Madrid 28006, Spain. xjuan@iqfr.csic.es.
Organizational Affiliation: