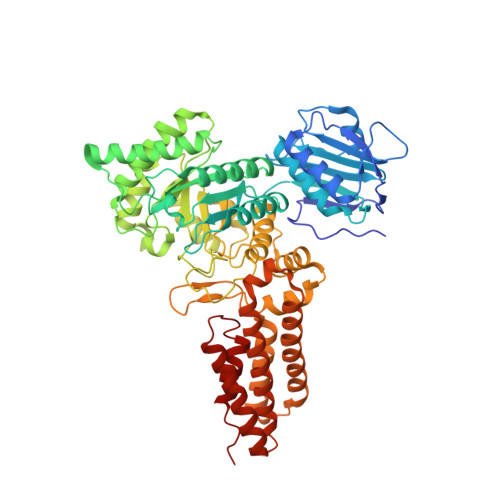
Glcnacstatins are Nanomolar Inhibitors of Human O-Glcnacase Inducing Cellular Hyper-O-Glcnacylation
Dorfmueller, H.C., Borodkin, V.S., Schimpl, M., Van Aalten, D.M.F.(2009) Biochem J 420: 221
- PubMed: 19275764 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1042/BJ20090110
- Primary Citation Related Structures:
2WB5 - PubMed Abstract:
O-GlcNAcylation is an essential, dynamic and inducible post-translational glycosylation of cytosolic proteins in metazoa and can show interplay with protein phosphorylation. Inhibition of OGA (O-GlcNAcase), the enzyme that removes O-GlcNAc from O-GlcNAcylated proteins, is a useful strategy to probe the role of this modification in a range of cellular processes. In the present study, we report the rational design and evaluation of GlcNAcstatins, a family of potent, competitive and selective inhibitors of human OGA. Kinetic experiments with recombinant human OGA reveal that the GlcNAcstatins are the most potent human OGA inhibitors reported to date, inhibiting the enzyme in the sub-nanomolar to nanomolar range. Modification of the GlcNAcstatin N-acetyl group leads to up to 160-fold selectivity against the human lysosomal hexosaminidases which employ a similar substrate-assisted catalytic mechanism. Mutagenesis studies in a bacterial OGA, guided by the structure of a GlcNAcstatin complex, provides insight into the role of conserved residues in the human OGA active site. GlcNAcstatins are cell-permeant and, at low nanomolar concentrations, effectively modulate intracellular O-GlcNAc levels through inhibition of OGA, in a range of human cell lines. Thus these compounds are potent selective tools to study the cell biology of O-GlcNAc.
- University of Dundee, Scotland, UK.
Organizational Affiliation: