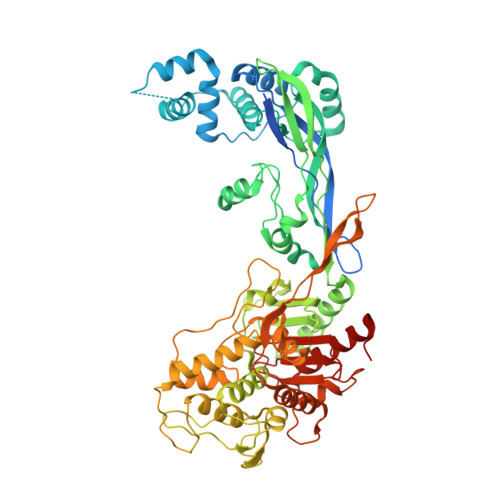
Pbp Active Site Flexibility as the Key Mechanism for Beta-Lactam Resistance in Pneumococci
Contreras-Martel, C., Dahout-Gonzalez, C., Dos-Santos-Martins, A., Kotnik, M., Dessen, A.(2009) J Mol Biology 387: 899
- PubMed: 19233207 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2009.02.024
- Primary Citation Related Structures:
2WAD, 2WAE, 2WAF - PubMed Abstract:
Penicillin-binding proteins (PBPs), the main targets of beta-lactam antibiotics, are membrane-associated enzymes that catalyze the two last steps in the biosynthesis of peptidoglycan. In Streptococcus pneumoniae, a major human pathogen, the surge in resistance to such antibiotics is a direct consequence of the proliferation of mosaic PBP-encoding genes, which give rise to proteins containing tens of mutations. PBP2b is a major drug resistance target, and its modification is essential for the development of high levels of resistance to piperacillin. In this work, we have solved the crystal structures of PBP2b from a wild-type pneumococcal strain, as well as from a highly drug-resistant clinical isolate displaying 58 mutations. Although mutations are present throughout the entire PBP structure, those surrounding the active site influence the total charge and the polar character of the region, while those in close proximity to the catalytic nucleophile impart flexibility onto the beta3/beta4 loop area, which encapsulates the cleft. The wealth of structural data on pneumococcal PBPs now underlines the importance of high malleability in active site regions of drug-resistant strains, suggesting that active site "breathing" could be a common mechanism employed by this pathogen to prevent targeting by beta-lactams.
- Institut de Biologie Structurale Jean-Pierre Ebel, UMR 5075 (CEA, CNRS, UJF, PSB), Grenoble, France.
Organizational Affiliation: