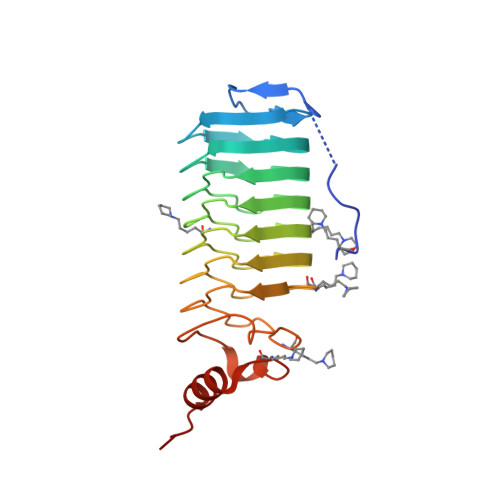
Crystallization of a Pentapeptide-Repeat Protein by Reductive Cyclic Pentylation of Free Amines with Glutaraldehyde.
Vetting, M.W., Hegde, S.S., Blanchard, J.S.(2009) Acta Crystallogr D Biol Crystallogr 65: 462
- PubMed: 19390151 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S0907444909008324
- Primary Citation Related Structures:
2W7Z - PubMed Abstract:
The pentapeptide-repeat protein EfsQnr from Enterococcus faecalis protects DNA gyrase from inhibition by fluoroquinolones. EfsQnr was cloned and purified to homogeneity, but failed to produce diffraction-quality crystals in initial crystallization screens. Treatment of EfsQnr with glutaraldehyde and the strong reducing agent borane-dimethylamine resulted in a derivatized protein which produced crystals that diffracted to 1.6 A resolution; their structure was subsequently determined by single-wavelength anomalous dispersion. Analysis of the derivatized protein using Fourier transform ion cyclotron resonance mass spectrometry indicated a mass increase of 68 Da per free amino group. Electron-density maps about a limited number of structurally ordered lysines indicated that the modification was a cyclic pentylation of free amines, producing piperidine groups.
- Department of Biochemistry, Albert Einstein College of Medicine, 1300 Morris Park Avenue, Bronx, NY 10461, USA. vetting@aecom.yu.edu
Organizational Affiliation: