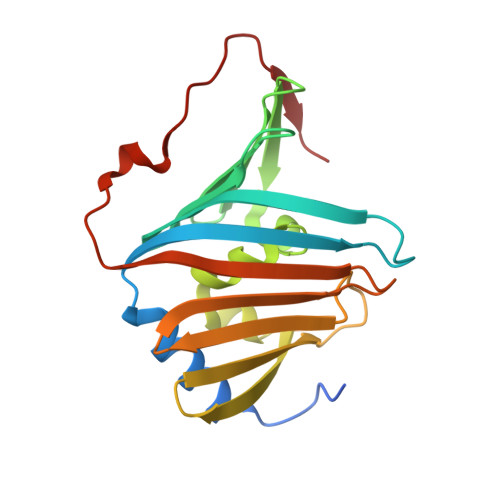
Hydrophobic Surface Patches on Lola of Pseudomonas Aeruginosa are Essential for Lipoprotein Binding.
Remans, K., Pauwels, K., Van Ulsen, P., Buts, L., Cornelis, P., Tommassen, J., Savvides, S., Decanniere, K., Van Gelder, P.(2010) J Mol Biology 401: 921
- PubMed: 20620146
- DOI: https://doi.org/10.1016/j.jmb.2010.06.067
- Primary Citation Related Structures:
2W7Q - PubMed Abstract:
Many lipoproteins reside in the outer membrane (OM) of Gram-negative bacteria, and their biogenesis is dependent on the Lol (localization of lipoproteins) system. The periplasmic chaperone LolA accepts OM-destined lipoproteins that are released from the inner membrane by the LolCDE complex and transfers them to the OM receptor LolB. The exact nature of the LolA-lipoprotein complex is still unknown. The crystal structure of Escherichia coli LolA features an open beta-barrel covered by alpha helices that together constitute a hydrophobic cavity, which would allow the binding of one acyl chain. However, OM lipoproteins contain three acyl chains, and the stoichiometry of the LolA-lipoprotein complex is 1:1. Here we present the crystal structure of Pseudomonas aeruginosa LolA that projects clear hydrophobic surface patches. Since these patches are large enough to accommodate acyl chains, their role in lipoprotein binding was investigated. Several LolA mutant proteins were created, and their functionality was assessed by studying their capacity to release lipoproteins produced in sphaeroplasts. Interruption of the largest hydrophobic patch completely destroyed the lipoprotein-releasing capacity of LolA, while interruption of smaller patches apparently reduced efficiency. Thus, the results show a new lipoprotein transport model that places (some of) the acyl chains on the hydrophobic surface patches.
- Department of Molecular and Cellular Interactions, Vrije Universiteit Brussel and VIB, 1050 Brussels, Belgium. kim.remans@vub.ac.be
Organizational Affiliation: