Structure and pathogenicity of antibodies specific for citrullinated collagen type II in experimental arthritis.
Uysal, H., Bockermann, R., Nandakumar, K.S., Sehnert, B., Bajtner, E., Engstrom, A., Serre, G., Burkhardt, H., Thunnissen, M.M., Holmdahl, R.(2009) J Exp Medicine 206: 449-462
- PubMed: 19204106 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1084/jem.20081862
- Primary Citation Related Structures:
2W60, 2W65 - PubMed Abstract:
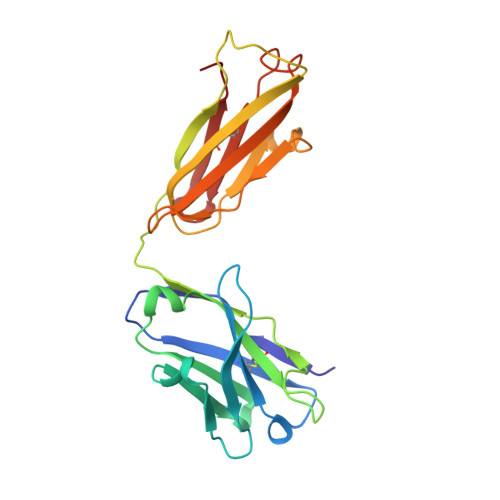
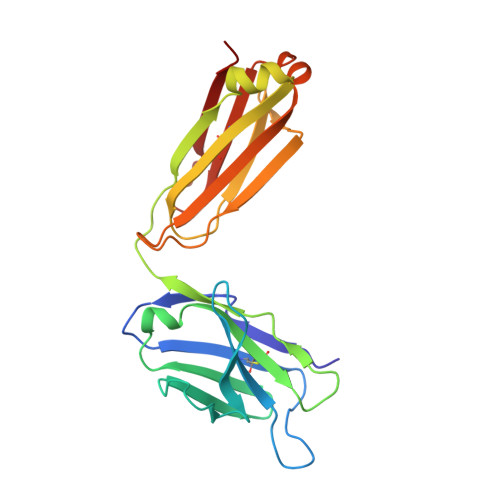
Antibodies to citrulline-modified proteins have a high diagnostic value in rheumatoid arthritis (RA). However, their biological role in disease development is still unclear. To obtain insight into this question, a panel of mouse monoclonal antibodies was generated against a major triple helical collagen type II (CII) epitope (position 359-369; ARGLTGRPGDA) with or without arginines modified by citrullination. These antibodies bind cartilage and synovial tissue, and mediate arthritis in mice. Detection of citrullinated CII from RA patients' synovial fluid demonstrates that cartilage-derived CII is indeed citrullinated in vivo. The structure determination of a Fab fragment of one of these antibodies in complex with a citrullinated peptide showed a surprising beta-turn conformation of the peptide and provided information on citrulline recognition. Based on these findings, we propose that autoimmunity to CII, leading to the production of antibodies specific for both native and citrullinated CII, is an important pathogenic factor in the development of RA.
- Center for Molecular Protein Science, Biomedical Center I11, Lund University, 221 00 Lund, Sweden.
Organizational Affiliation: