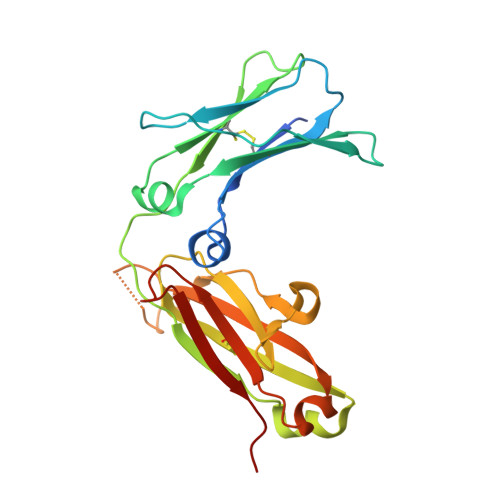
The Crystal Structure of an Avian Igy-Fc Fragment Reveals Conservation with Both Mammalian Igg and Ige.
Taylor, A.I., Fabiane, S.M., Sutton, B.J., Calvert, R.A.(2009) Biochemistry 48: 558
- PubMed: 19115948 Search on PubMed
- DOI: https://doi.org/10.1021/bi8019993
- Primary Citation Related Structures:
2W59 - PubMed Abstract:
Avian IgY is closely related to an ancestor of both mammalian IgG and IgE and thus provides insights into the evolution of antibody structure and function. A recombinant fragment of IgY-Fc consisting of a dimer of the Cupsilon3 and Cupsilon4 domains, Fcupsilon3-4, was expressed and crystallized and its X-ray structure determined to 1.75 A resolution. Fcupsilon3-4 is the only nonmammalian Fc fragment structure determined to date and provides the first structural evidence for an ancient origin of antibody architecture. The Fcupsilon3-4 structure reveals features common to both IgE-Fc and IgG-Fc, and the implications for IgY binding to its receptor are discussed.
- Randall Division of Cell and Molecular Biophysics, King's College London, and MRC-Asthma UK Centre in Allergic Mechanisms of Asthma, New Hunt's House, Guy's Campus, London SE1 1UL, United Kingdom.
Organizational Affiliation: