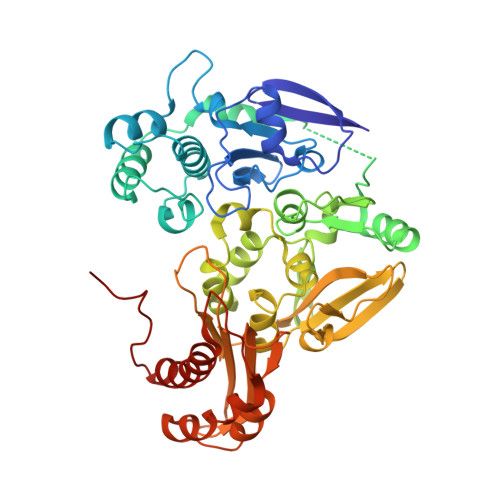
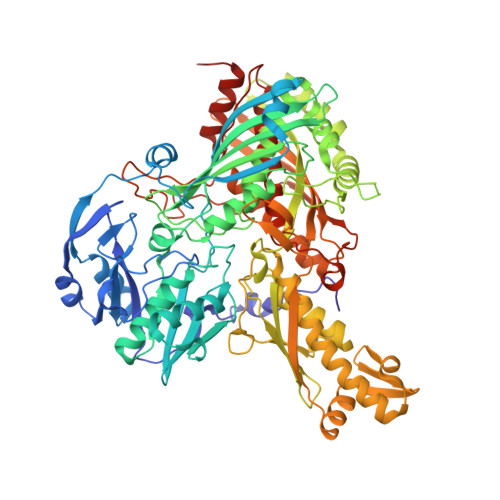
Mechanism of Substrate and Inhibitor Binding of Rhodobacter Capsulatus Xanthine Dehydrogenase.
Dietzel, U., Kuper, J., Doebbler, J.A., Schulte, A., Truglio, J.J., Leimkuhler, S., Kisker, C.(2009) J Biological Chem 284: 8768
- PubMed: 19109249 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M808114200
- Primary Citation Related Structures:
2W3R, 2W3S, 2W54, 2W55 - PubMed Abstract:
Rhodobacter capsulatus xanthine dehydrogenase (XDH) is an (alphabeta)(2) heterotetrameric cytoplasmic enzyme that resembles eukaryotic xanthine oxidoreductases in respect to both amino acid sequence and structural fold. To obtain a detailed understanding of the mechanism of substrate and inhibitor binding at the active site, we solved crystal structures of R. capsulatus XDH in the presence of its substrates hypoxanthine, xanthine, and the inhibitor pterin-6-aldehyde using either the inactive desulfo form of the enzyme or an active site mutant (E(B)232Q) to prevent substrate turnover. The hypoxanthine- and xanthine-bound structures reveal the orientation of both substrates at the active site and show the importance of residue Glu(B)-232 for substrate positioning. The oxygen atom at the C-6 position of both substrates is oriented toward Arg(B)-310 in the active site. Thus the substrates bind in an orientation opposite to the one seen in the structure of the reduced enzyme with the inhibitor oxypurinol. The tightness of the substrates in the active site suggests that the intermediate products must exit the binding pocket to allow first the attack of the C-2, followed by oxidation of the C-8 atom to form the final product uric acid. Structural studies of pterin-6-aldehyde, a potent inhibitor of R. capsulatus XDH, contribute further to the understanding of the relative positioning of inhibitors and substrates in the binding pocket. Steady state kinetics reveal a competitive inhibition pattern with a K(i) of 103.57 +/- 18.96 nm for pterin-6-aldehyde.
- Rudolf Virchow Center for Experimental Biomedicine, Institute for Structural Biology, University of Würzburg, Versbacher Strasse 9, 97078 Würzburg, Germany.
Organizational Affiliation: