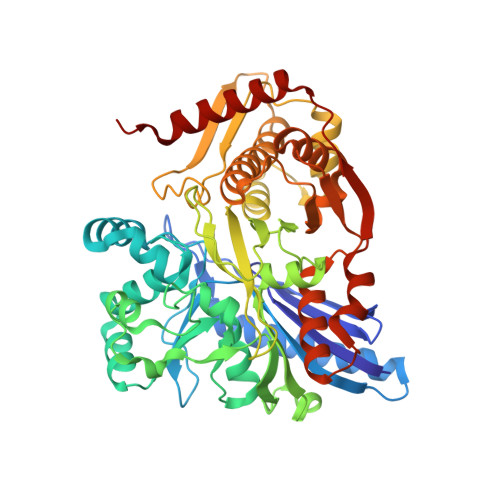
Structure and Non-Essential Function of Glycerol Kinase in Plasmodium Falciparum Blood Stages.
Schnick, C., Polley, S.D., Fivelman, Q.L., Ranford-Cartwright, L., Wilkinson, S.R., Brannigan, J.A., Wilkinson, A.J., Baker, D.A.(2009) Mol Microbiol 71: 533
- PubMed: 19040641 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1111/j.1365-2958.2008.06544.x
- Primary Citation Related Structures:
2W40, 2W41 - PubMed Abstract:
Malaria pathology is caused by multiplication of asexual parasites within erythrocytes, whereas mosquito transmission of malaria is mediated by sexual precursor cells (gametocytes). Microarray analysis identified glycerol kinase (GK) as the second most highly upregulated gene in Plasmodium falciparum gametocytes with no expression detectable in asexual blood stage parasites. Phosphorylation of glycerol by GK is the rate-limiting step in glycerol utilization. Deletion of this gene from P. falciparum had no effect on asexual parasite growth, but surprisingly also had no effect on gametocyte development or exflagellation, suggesting that these life cycle stages do not utilize host-derived glycerol as a carbon source. Kinetic studies of purified PfGK showed that the enzyme is not regulated by fructose 1,6 bisphosphate. The high-resolution crystal structure of P. falciparum GK, the first of a eukaryotic GK, reveals two domains embracing a capacious ligand-binding groove. In the complexes of PfGK with glycerol and ADP, we observed closed and open forms of the active site respectively. The 27 degree domain opening is larger than in orthologous systems and exposes an extensive surface with potential for exploitation in selective inhibitor design should the enzyme prove to be essential in vivo either in the human or in the mosquito.
- Structural Biology Laboratory, Department of Chemistry, University of York, York YO10 5YW, UK.
Organizational Affiliation: