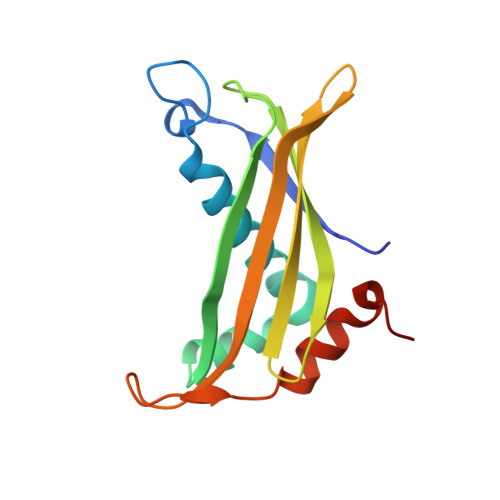
Structure and Catalytic Mechanism of the Thioesterase Cale7 in Enediyne Biosynthesis.
Kotaka, M., Kong, R., Qureshi, I., Ho, Q.S., Sun, H., Liew, C.W., Goh, L.P., Cheung, P., Mu, Y., Lescar, J., Liang, Z.X.(2009) J Biological Chem 284: 15739
- PubMed: 19357082 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M809669200
- Primary Citation Related Structures:
2W3X - PubMed Abstract:
The biosynthesis of the enediyne moiety of the antitumor natural product calicheamicin involves an iterative polyketide synthase (CalE8) and other ancillary enzymes. In the proposed mechanism for the early stage of 10-membered enediyne biosynthesis, CalE8 produces a carbonyl-conjugated polyene with the assistance of a putative thioesterase (CalE7). We have determined the x-ray crystal structure of CalE7 and found that the subunit adopts a hotdog fold with an elongated and kinked substrate-binding channel embedded between two subunits. The 1.75-A crystal structure revealed that CalE7 does not contain a critical catalytic residue (Glu or Asp) conserved in other hotdog fold thioesterases. Based on biochemical and site-directed mutagenesis studies, we proposed a catalytic mechanism in which the conserved Arg(37) plays a crucial role in the hydrolysis of the thioester bond, and that Tyr(29) and a hydrogen-bonded water network assist the decarboxylation of the beta-ketocarboxylic acid intermediate. Moreover, computational docking suggested that the substrate-binding channel binds a polyene substrate that contains a single cis double bond at the C4/C5 position, raising the possibility that the C4=C5 double bond in the enediyne moiety could be generated by the iterative polyketide synthase. Together, the results revealed a hotdog fold thioesterase distinct from the common type I and type II thioesterases associated with polyketide biosynthesis and provided interesting insight into the enediyne biosynthetic mechanism.
- School of Biological Sciences, Nanyang Technological University, 60 Nanyang Drive, Singapore 637551, Singapore.
Organizational Affiliation: