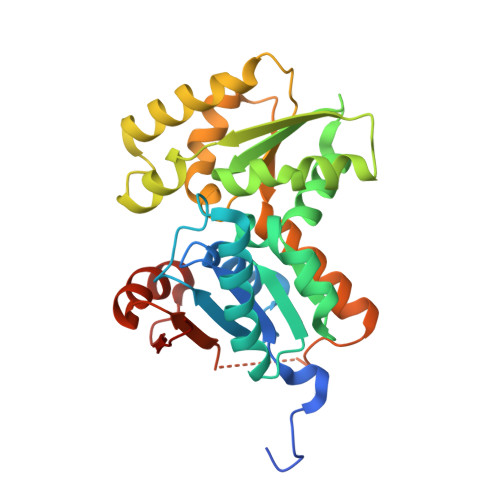
Exploring 9-Benzyl Purines as Inhibitors of Glutamate Racemase (Muri) in Gram-Positive Bacteria.
Geng, B., Breault, G., Comita-Prevoir, J., Petrichko, R., Eyermann, C., Lundqvist, T., Doig, P., Gorseth, E., Noonan, B.(2008) Bioorg Med Chem Lett 18: 4368
- PubMed: 18614367 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2008.06.068
- Primary Citation Related Structures:
2VVT - PubMed Abstract:
An early SAR study of a screening hit series has generated a series of 9-benzyl purines as inhibitors of bacterial glutamate racemase (MurI) with micromolar enzyme potency and improved physical properties. X-ray co-crystal EI structures were obtained.
- AstraZeneca R&D Boston, Infection Discovery, 35 Gatehouse Drive, Waltham, MA 02451, USA. bolin.geng@astrazeneca.com
Organizational Affiliation: