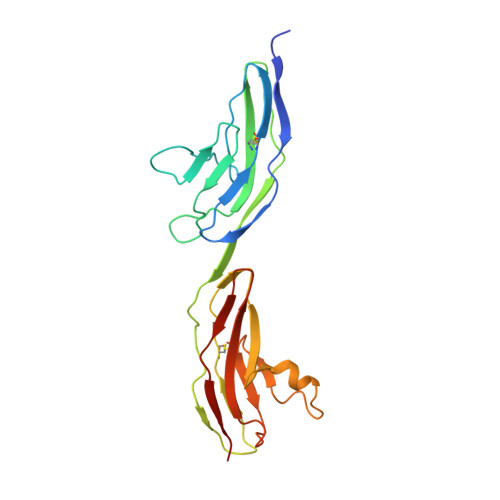
Structural and Functional Analysis of Slit and Heparin Binding to Immunoglobulin-Like Domains 1 and 2 of Drosophila Robo
Fukuhara, N., Howitt, J.A., Hussain, S., Hohenester, E.(2008) J Biological Chem 283: 16226
- PubMed: 18359766 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M800688200
- Primary Citation Related Structures:
2VR9, 2VRA - PubMed Abstract:
Recognition of the secreted protein Slit by transmembrane receptors of the Robo family provides important signals in the development of the nervous system and other organs, as well as in tumor metastasis and angiogenesis. Heparan sulfate (HS) proteoglycans serve as essential co-receptors in Slit-Robo signaling. Previous studies have shown that the second leucinerich repeat domain of Slit, D2, binds to the N-terminal immunoglobulin-like domains of Robo, IG1-2. Here we present two crystal structures of Drosophila Robo IG1-2, one of which contains a bound heparin-derived oligosaccharide. Using structure-based mutagenesis of a Robo IG1-5 construct we identified key Slit binding residues (Thr-74, Phe-114, Arg-117) forming a conserved patch on the surface of IG1; mutation of similarly conserved residues in IG2 had no effect on Slit binding. Mutation of conserved basic residues in IG1 (Lys-69, Arg-117, Lys-122, Lys-123), but not in IG2, reduced binding of Robo IG1-5 to heparin, in full agreement with the Robo-heparin co-crystal structure. Our collective results, together with a recent crystal structure of a minimal human Slit-Robo complex ( Morlot, C., Thielens, N. M., Ravelli, R. B., Hemrika, W., Romijn, R. A., Gros, P., Cusack, S., and McCarthy, A. A. (2007) Proc. Natl. Acad. Sci. U.S.A. 104, 14923-14928 ), reveal a contiguous HS/heparin binding surface extending across the Slit-Robo interface. Based on the size of this composite binding site, we predict that at least five HS disaccharide units are required to support Slit-Robo signaling.
- Department of Life Sciences, Imperial College London, London SW7 2AZ, United Kingdom.
Organizational Affiliation: